
| Sequence ID | dm3.chr3LHet |
|---|---|
| Location | 1,315,179 – 1,315,283 |
| Length | 104 |
| Max. P | 0.586034 |

| Location | 1,315,179 – 1,315,283 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 84.05 |
| Shannon entropy | 0.20990 |
| G+C content | 0.50412 |
| Mean single sequence MFE | -26.91 |
| Consensus MFE | -24.00 |
| Energy contribution | -23.78 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.13 |
| Structure conservation index | 0.89 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.586034 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3LHet 1315179 104 + 2555491 CUAUAGGCCCAUAAAACUCUAUCCCCAGGCUCUCGAUAAGCGUAAACAAUCGGAGUGUCCUUGCCUGGGGCCAAUUCAGGAGUACGUU-UUCGAGCUCCAACUAA ...(((................((((((((....((((..((........))...))))...))))))))........(((((.((..-..)).)))))..))). ( -31.10, z-score = -2.05, R) >droSim1.chrU 13443535 91 - 15797150 CUAUAGGCCCAUAAAACUCUAUCCACAGGCUAUCGAUAAGUGC--------------CCCUUGCCUGGGGUCAAUUCAGGAGUACGUUGUCCGAGCUCCAUCUGA .....((((((((......)))...(((((.............--------------.....))))))))))......(((((.((.....)).)))))...... ( -23.47, z-score = -0.25, R) >droSec1.super_48 113846 91 + 230265 CUAUAGGCCCAUAAAACUCUAUCCCCAGGCUAUCGAUAAGUGC--------------CCCUUGCCUGGGGUCAAUUCAGGAGUACGUUGUCCGAGCUCCAUCUGA ...((((...............((((((((.............--------------.....))))))))........(((((.((.....)).))))).)))). ( -26.17, z-score = -1.09, R) >consensus CUAUAGGCCCAUAAAACUCUAUCCCCAGGCUAUCGAUAAGUGC______________CCCUUGCCUGGGGUCAAUUCAGGAGUACGUUGUCCGAGCUCCAUCUGA .....((((((((......)))...(((((................................))))))))))......(((((.((.....)).)))))...... (-24.00 = -23.78 + -0.22)

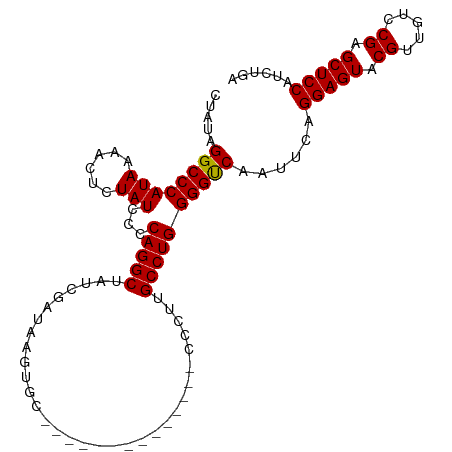
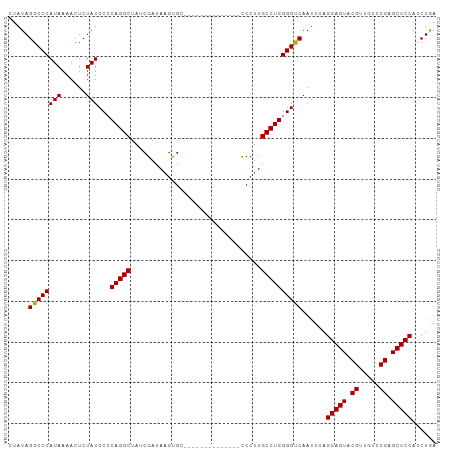
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:51:15 2011