
| Sequence ID | dm3.chr3LHet |
|---|---|
| Location | 517,378 – 517,479 |
| Length | 101 |
| Max. P | 0.991732 |

| Location | 517,378 – 517,479 |
|---|---|
| Length | 101 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 51.60 |
| Shannon entropy | 0.68599 |
| G+C content | 0.45220 |
| Mean single sequence MFE | -29.03 |
| Consensus MFE | -9.13 |
| Energy contribution | -9.37 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.59 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.18 |
| SVM RNA-class probability | 0.905945 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
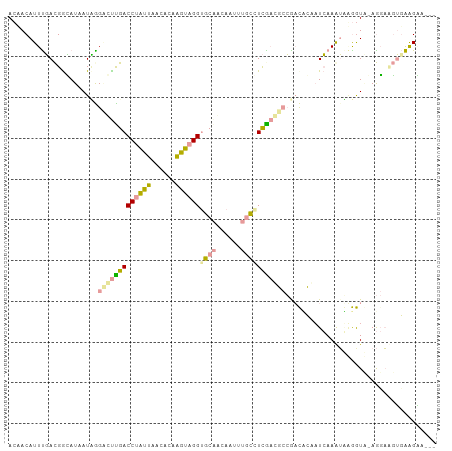
>dm3.chr3LHet 517378 101 + 2555491 ACAACAUUCGACGGUAUAAUCGGGGCCGACCUGUUAACACAGGCCGGUGCAUCACUUUGCCUCGCCUCCGCCCAACUCAAAUGGGGUC-AGGAAGUUGAGAA--- .((((.((((((((......((((((((.(((((....))))).))))(((......)))))))...)).((((.......)))))))-.))).))))....--- ( -33.10, z-score = -1.07, R) >droEre2.scaffold_4845 20482470 102 - 22589142 AAAACAUUUGACGGCAUAAUUAGCCUUGACCUAUUAGCGCAAGUAGGUGCGACAAUUGACGUCAAGGCCAAUUCCAUCAAGUAUAAUAGAGGACGUGAAGGA--- .....((((((.((...((((.((((((((..(((..((((......))))..)))....)))))))).)))))).))))))....................--- ( -25.70, z-score = -1.84, R) >droPer1.super_109 48952 105 - 183219 ACUACAUUUUAUGAGAUCAUAGGACUUGACCUACUAAAACAAGUAGGGGCAGUAUUAUGUCUUGAUGAAGACACAAUAAAAAUAGGUGCAACUUCGGAGGCGCUG ......(((((((....))))))).(((.((((((......))))))..)))((((.((((((....)))))).))))......(((((..(....)..))))). ( -28.30, z-score = -3.19, R) >consensus ACAACAUUUGACGGCAUAAUAGGACUUGACCUAUUAACACAAGUAGGUGCAACAAUUUGCCUCGACGCCGACACAAUCAAAUAAGGUA_AGGAAGUGAAGAA___ ......................(((((((((((((......))))))((((......)))))))))))..................................... ( -9.13 = -9.37 + 0.24)
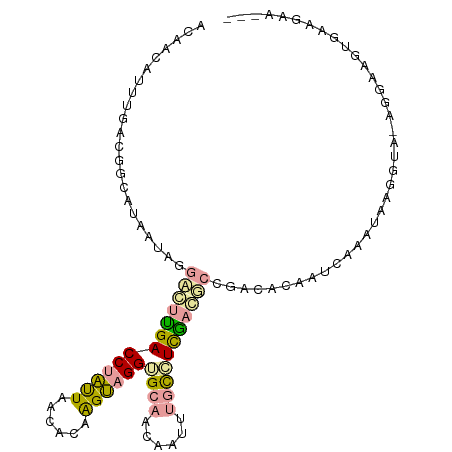


| Location | 517,378 – 517,479 |
|---|---|
| Length | 101 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 51.60 |
| Shannon entropy | 0.68599 |
| G+C content | 0.45220 |
| Mean single sequence MFE | -30.93 |
| Consensus MFE | -7.93 |
| Energy contribution | -7.40 |
| Covariance contribution | -0.53 |
| Combinations/Pair | 1.68 |
| Mean z-score | -3.08 |
| Structure conservation index | 0.26 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.49 |
| SVM RNA-class probability | 0.991732 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
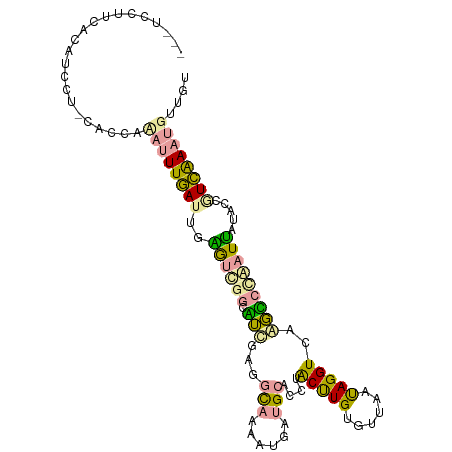
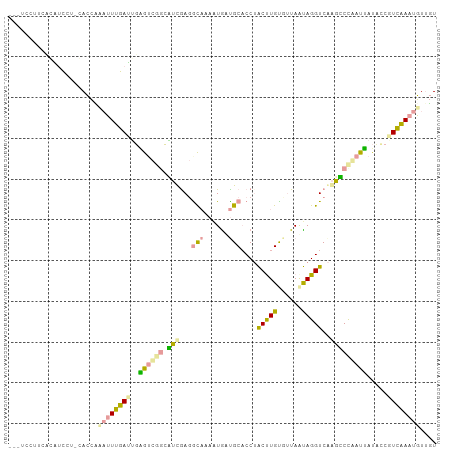
>dm3.chr3LHet 517378 101 - 2555491 ---UUCUCAACUUCCU-GACCCCAUUUGAGUUGGGCGGAGGCGAGGCAAAGUGAUGCACCGGCCUGUGUUAACAGGUCGGCCCCGAUUAUACCGUCGAAUGUUGU ---....(((((((..-(((((((.......))))(((.(((...(((......)))..((((((((....))))))))))))))........)))))).)))). ( -36.30, z-score = -1.68, R) >droEre2.scaffold_4845 20482470 102 + 22589142 ---UCCUUCACGUCCUCUAUUAUACUUGAUGGAAUUGGCCUUGACGUCAAUUGUCGCACCUACUUGCGCUAAUAGGUCAAGGCUAAUUAUGCCGUCAAAUGUUUU ---..................(((.((((((((((((((((((((.(..((((.((((......)))).))))).)))))))))))))...))))))).)))... ( -32.50, z-score = -4.67, R) >droPer1.super_109 48952 105 + 183219 CAGCGCCUCCGAAGUUGCACCUAUUUUUAUUGUGUCUUCAUCAAGACAUAAUACUGCCCCUACUUGUUUUAGUAGGUCAAGUCCUAUGAUCUCAUAAAAUGUAGU ..(((.((....)).))).........(((((((((((....)))))))))))((((.((((((......))))))........((((....))))....)))). ( -24.00, z-score = -2.90, R) >consensus ___UCCUUCACAUCCU_CACCAAAUUUGAUUGAGUCGGCAUCGAGGCAAAAUGAUGCACCUACUUGUGUUAAUAGGUCAAGCCCAAUUAUACCGUCAAAUGUUGU ......................((((((((..((((((.(((...(((......)))....(((((......)))))..))))))))).....)))))))).... ( -7.93 = -7.40 + -0.53)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:50:39 2011