
| Sequence ID | dm3.chr3LHet |
|---|---|
| Location | 500,139 – 500,256 |
| Length | 117 |
| Max. P | 0.836737 |

| Location | 500,139 – 500,253 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 53.51 |
| Shannon entropy | 0.84726 |
| G+C content | 0.43684 |
| Mean single sequence MFE | -33.87 |
| Consensus MFE | -7.70 |
| Energy contribution | -8.06 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.87 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.23 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.836737 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3LHet 500139 114 - 2555491 CUAGAACUCUGCUAACAAAGGUGACAUCAAUGAAAGAUGUACCUCUGUCAUUAUUGAACGUCGGUUUGAACCCGUCGUUUAGCAAUGCGAGAUCUAGCAGGUUCAUGGUAUCGA (((((((.(((((((((.(((((.((((.......))))))))).)))((((.(((((((.(((.......))).))))))).)))).......)))))))))).)))...... ( -37.60, z-score = -3.42, R) >dp4.chrXR_group5 67475 114 - 740492 CUAGUUGAGUACUUAUGUAGGUGAGGUCUAUGAUGGAUGUGGUAUUUGCUCUGCUGAAAGUUGGGUUGCAACCAGCGUUUUUUACGACUAUGUCCAGGGAGGCAAAAGCCUCCA .......((..(((((....)))))..))....((((((((((........(((((...((((.....))))))))).........)))))))))).((((((....)))))). ( -34.13, z-score = -0.96, R) >droPer1.super_166 55330 114 + 96193 CUAGUCGAGUACUUAUGUAGGUGAGGUCUAUGAUGGCUGUGGUACCUGCUCUGCUGAAUGUUGGGUUGGAACCAGCGUUUUUUAGGACUAUGUCCAGGGAGGCAAAAGCCUCCA .(((((..........(((((((.((((......))))....))))))).(((..(((((((((.......)))))))))..)))))))).......((((((....)))))). ( -43.80, z-score = -2.89, R) >droWil1.scaffold_181145 347793 114 - 2682060 CCAGGCUCGUGCUUGAAAAGGUAACAUCAAUUUUUGAGUUACGUCCUGCUUUGCUGAAUGUUUGUGUGCUGCCGAUAUUCAGUAAACUCACGUUUCUCGAGGCAAAGGAGUCCA ...(((((.(((((....)))))........(((((((..((((...(.(((((((((((((.((.....)).))))))))))))))..))))..))))))).....))))).. ( -35.80, z-score = -2.42, R) >triCas2.ChLG10 1594212 114 - 8806720 CAUUAUUUUUUCAGACACUAGUGAGGUCAAUGUAUAAUGAUCCGUUUGUGCGCUCUAGUGUAGGUACUUUACCGUCAUUUAAUACGCUCAUAUUCAUAAAUUCUGCAAGUUCCA .......(((.((((.....(((((.....(((((((((((.......(((((....)))))((((...))))))))))..)))))......)))))....)))).)))..... ( -18.00, z-score = 0.36, R) >consensus CUAGACGAGUACUUACAAAGGUGAGGUCAAUGAUGGAUGUACCACUUGCUCUGCUGAAUGUUGGUUUGCAACCGGCGUUUAGUAAGACCAUGUCCAGCGAGGCAAAAGCCUCCA .........(((((....))))).........................((((((((((((((((.......)))))))))............((....)))))))))....... ( -7.70 = -8.06 + 0.36)



| Location | 500,142 – 500,256 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 52.02 |
| Shannon entropy | 0.87700 |
| G+C content | 0.43684 |
| Mean single sequence MFE | -32.21 |
| Consensus MFE | -7.70 |
| Energy contribution | -8.06 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.87 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.24 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr3LHet 500142 114 - 2555491 CCACUAGAACUCUGCUAACAAAGGUGACAUCAAUGAAAGAUGUACCUCUGUCAUUAUUGAACGUCGGUUUGAACCCGUCGUUUAGCAAUGCGAGAUCUAGCAGGUUCAUGGUAU ..((((((((.(((((((((.(((((.((((.......))))))))).)))((((.(((((((.(((.......))).))))))).)))).......)))))))))).)))).. ( -39.70, z-score = -4.34, R) >dp4.chrXR_group5 67478 114 - 740492 CUGCUAGUUGAGUACUUAUGUAGGUGAGGUCUAUGAUGGAUGUGGUAUUUGCUCUGCUGAAAGUUGGGUUGCAACCAGCGUUUUUUACGACUAUGUCCAGGGAGGCAAAAGCCU .(((.(((.....)))...)))(((...((((....((((((((((........(((((...((((.....))))))))).........))))))))))...))))....))). ( -30.13, z-score = 0.24, R) >droPer1.super_166 55333 114 + 96193 CUGCUAGUCGAGUACUUAUGUAGGUGAGGUCUAUGAUGGCUGUGGUACCUGCUCUGCUGAAUGUUGGGUUGGAACCAGCGUUUUUUAGGACUAUGUCCAGGGAGGCAAAAGCCU (((.(((((..........(((((((.((((......))))....))))))).(((..(((((((((.......)))))))))..))))))))....)))..((((....)))) ( -37.60, z-score = -1.17, R) >droWil1.scaffold_181145 347796 114 - 2682060 GGGCCAGGCUCGUGCUUGAAAAGGUAACAUCAAUUUUUGAGUUACGUCCUGCUUUGCUGAAUGUUUGUGUGCUGCCGAUAUUCAGUAAACUCACGUUUCUCGAGGCAAAGGAGU (((((((((....))))).....(((((.(((.....))))))))))))(((((((..((((((..((.(((((........))))).))..))))))..)))))))....... ( -35.60, z-score = -1.70, R) >triCas2.ChLG10 1594215 114 - 8806720 UUCCAUUAUUUUUUCAGACACUAGUGAGGUCAAUGUAUAAUGAUCCGUUUGUGCGCUCUAGUGUAGGUACUUUACCGUCAUUUAAUACGCUCAUAUUCAUAAAUUCUGCAAGUU ..........(((.((((.....(((((.....(((((((((((.......(((((....)))))((((...))))))))))..)))))......)))))....)))).))).. ( -18.00, z-score = 0.28, R) >consensus CUGCUAGACGAGUACUUACAAAGGUGAGGUCAAUGAUGGAUGUACCACUUGCUCUGCUGAAUGUUGGUUUGCAACCGGCGUUUAGUAAGACCAUGUCCAGCGAGGCAAAAGCCU ............(((((....))))).........................((((((((((((((((.......)))))))))............((....))))))))).... ( -7.70 = -8.06 + 0.36)


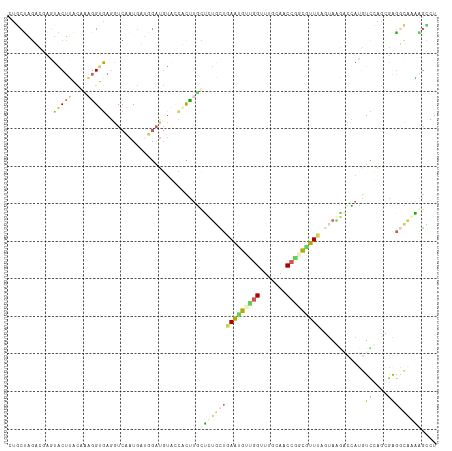
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:50:36 2011