| Sequence ID | dm3.chr3LHet |
|---|---|
| Location | 487,516 – 487,608 |
| Length | 92 |
| Max. P | 0.999766 |

| Location | 487,516 – 487,608 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 55.13 |
| Shannon entropy | 0.62791 |
| G+C content | 0.48591 |
| Mean single sequence MFE | -31.77 |
| Consensus MFE | -15.04 |
| Energy contribution | -16.50 |
| Covariance contribution | 1.46 |
| Combinations/Pair | 1.46 |
| Mean z-score | -3.32 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 4.34 |
| SVM RNA-class probability | 0.999766 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3LHet 487516 92 - 2555491 -----------UACAAAGAGGAUAACUGGAGGCGCUUAGUGGGAGAGAAUAAGGACGAUCCAUGGGGGCAAGUCUUUAGAAUUUUCCGUGGCCGCAAAAGGAC---- -----------..............((....(((((...(((((((...(((((((..(((....)))...)))))))...))))))).)).)))....))..---- ( -23.40, z-score = -1.23, R) >droAna3.scaffold_13266 19317466 97 - 19884421 ---------GAGAGAGAGAAGAUGGGUGGCGCAUUUUCGUAGAACAAAAUAAGCACGAUCCUUGGGGUCGUGUUUACAAAGUUCUGCGGGAGAGUGGUA-AGAAUUG ---------...........(((...((.(((.((..((((((((....((((((((((((...))))))))))))....))))))))..)).))).))-...))). ( -36.00, z-score = -5.85, R) >droGri2.scaffold_15040 118951 107 - 162499 UGCUUGUAAGAGUGAAAGAGGAUGAAUGGCGUUCCUUUUUGGAGCGCAAUAAGGACGACCCCUGGGGUCGUGCUUAUAAAGUUGUGAGAGGUAGGCGUAGGGAAGCG .((((.(....((...............(((((((.....))))))).(((((.(((((((...))))))).))))).................))....).)))). ( -35.90, z-score = -2.89, R) >consensus _________GAGAGAAAGAGGAUGAAUGGCGCACCUUAGUGGAACAAAAUAAGGACGAUCCAUGGGGUCGUGUUUAUAAAGUUCUGCGAGGCAGCAGUAGGGAA__G .................................((((((((((((...(((((.(((((((...))))))).)))))...))))))))))))............... (-15.04 = -16.50 + 1.46)
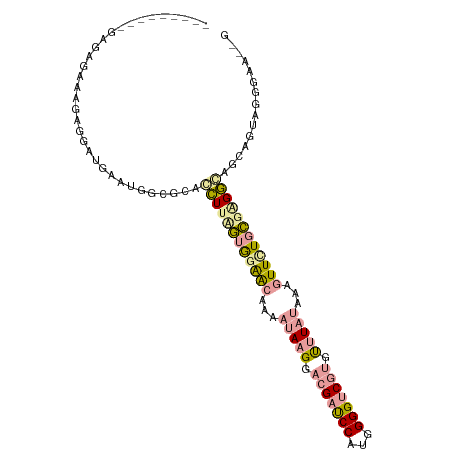


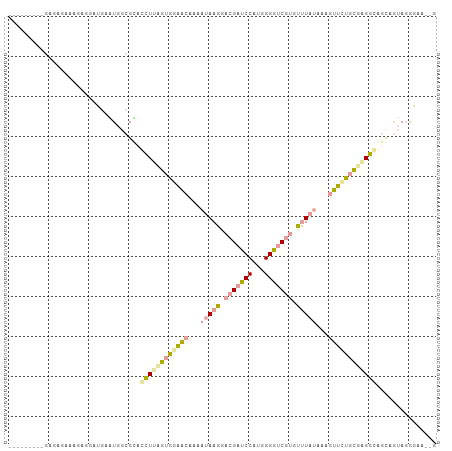
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:50:34 2011