
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 3,934,857 – 3,934,949 |
| Length | 92 |
| Max. P | 0.792060 |

| Location | 3,934,857 – 3,934,948 |
|---|---|
| Length | 91 |
| Sequences | 8 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 80.27 |
| Shannon entropy | 0.38410 |
| G+C content | 0.51387 |
| Mean single sequence MFE | -26.94 |
| Consensus MFE | -16.35 |
| Energy contribution | -17.49 |
| Covariance contribution | 1.14 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.768756 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
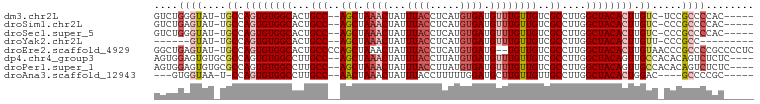
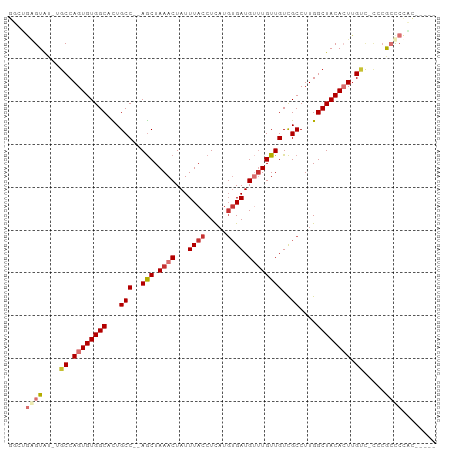
>dm3.chr2L 3934857 91 + 23011544 GUCUGGGUAU-UGCCAGUGUGGCACUGCC--AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUC-UCCGCCCCAC----- ....((((..-.((.((((((((...(((--(((.((((...((((.....)))).))))))))..))....)))))))).)).-...))))...----- ( -26.50, z-score = -1.47, R) >droSim1.chr2L 3889851 91 + 22036055 GUCUGAGUAU-UGCCAGUGUGGCACUGCC--AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUC-CCCGCCCCAC----- ..........-.((.((((((((...(((--(((.((((...((((.....)))).))))))))..))....)))))))).)).-..........----- ( -20.50, z-score = -0.23, R) >droSec1.super_5 2037832 91 + 5866729 GUCUGGGUAU-UGCCAGUGUGGCACUGCC--AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUC-CCCGCCCCAC----- ....((((..-.((.((((((((...(((--(((.((((...((((.....)))).))))))))..))....)))))))).)).-...))))...----- ( -26.50, z-score = -1.37, R) >droYak2.chr2L 3949134 81 + 22324452 ------GUAU-UGCCAGUGUGGCACUGCC--AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUU-CCCGCC--------- ------((..-.((.((((((((...(((--(((.((((...((((.....)))).))))))))..))....)))))))).)).-...)).--------- ( -20.60, z-score = -1.14, R) >droEre2.scaffold_4929 3992729 97 + 26641161 GGCUGAGUAU-UGCCAGUGUGGCACUGCCCCAGCUAAACUAUUUACCUCAUGUGAUG--UGUUGUCGCCUUGGCUACACUUGUAACCCGCCCCGCCCCUC (((.(.((.(-(((.((((((((...((..((((...((...((((.....)))).)--)))))..))....)))))))).))))...)).).))).... ( -26.50, z-score = -1.34, R) >dp4.chr4_group3 9602991 94 - 11692001 AGUGGAGUGUGCGCCAGUGUGGCCUUGCC--AGCUAAACUAUUUACCUUAUGUGAUGUUUGUUGUCGCCUUGGCUACAGUUGCCACACAGUCUCUC---- ......(((((.((.(.(((((((..(((--(((.((((...((((.....)))).))))))))..))...))))))).).)))))))........---- ( -31.00, z-score = -1.77, R) >droPer1.super_1 6717849 94 - 10282868 AGUGGAGUGUGCGCCAGUGUGGCCUUGCC--AGCUAAACUAUUUACCUUAUGUGAUGUUUGUUGUCGCCUUGGCUACAGUUGCCACACAGUCUCUC---- ......(((((.((.(.(((((((..(((--(((.((((...((((.....)))).))))))))..))...))))))).).)))))))........---- ( -31.00, z-score = -1.77, R) >droAna3.scaffold_12943 481321 84 - 5039921 ---GUGGUAA-U-CCAGUGUGGCCUUGCC--AACUAAACUAUUUACCUUUUUGGAUGCUUGUUGUUGCCUUGGCUACACUGGAC----GCCCCGC----- ---((((...-(-(((((((((((..(.(--(((..((.((((((......)))))).))...)))).)..)))))))))))).----...))))----- ( -32.90, z-score = -4.74, R) >consensus GGCUGAGUAU_UGCCAGUGUGGCACUGCC__AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUC_CCCGCCCCAC_____ ....((((....((.((((((((...((...(((.((((...((((.....)))).)))))))...))....)))))))).)).....))))........ (-16.35 = -17.49 + 1.14)

| Location | 3,934,858 – 3,934,949 |
|---|---|
| Length | 91 |
| Sequences | 8 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 80.53 |
| Shannon entropy | 0.37816 |
| G+C content | 0.51449 |
| Mean single sequence MFE | -26.40 |
| Consensus MFE | -16.35 |
| Energy contribution | -17.49 |
| Covariance contribution | 1.14 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.792060 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
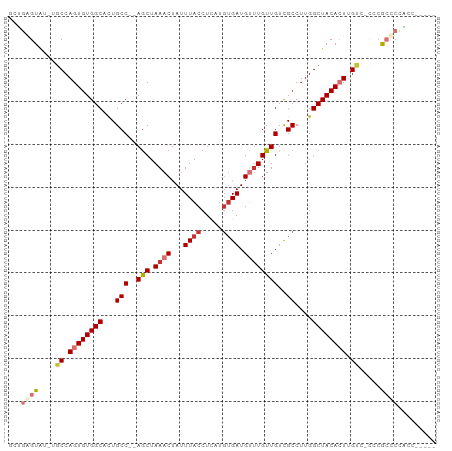
>dm3.chr2L 3934858 91 + 23011544 UCUGGGUAU-UGCCAGUGUGGCACUGCC--AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUC-UCCGCCCCACC----- ...((((..-.((.((((((((...(((--(((.((((...((((.....)))).))))))))..))....)))))))).)).-...))))....----- ( -26.50, z-score = -1.67, R) >droSim1.chr2L 3889852 91 + 22036055 UCUGAGUAU-UGCCAGUGUGGCACUGCC--AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUC-CCCGCCCCACC----- .........-.((.((((((((...(((--(((.((((...((((.....)))).))))))))..))....)))))))).)).-...........----- ( -20.50, z-score = -0.51, R) >droSec1.super_5 2037833 91 + 5866729 UCUGGGUAU-UGCCAGUGUGGCACUGCC--AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUC-CCCGCCCCACC----- ...((((..-.((.((((((((...(((--(((.((((...((((.....)))).))))))))..))....)))))))).)).-...))))....----- ( -26.50, z-score = -1.58, R) >droYak2.chr2L 3949134 81 + 22324452 -----GUAU-UGCCAGUGUGGCACUGCC--AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUU-CCCGCC---------- -----((..-.((.((((((((...(((--(((.((((...((((.....)))).))))))))..))....)))))))).)).-...)).---------- ( -20.60, z-score = -1.14, R) >droEre2.scaffold_4929 3992730 97 + 26641161 GCUGAGUAU-UGCCAGUGUGGCACUGCCCCAGCUAAACUAUUUACCUCAUGUGAUG--UGUUGUCGCCUUGGCUACACUUGUAACCCGCCCCGCCCCUCA ((.(.((.(-(((.((((((((...((..((((...((...((((.....)))).)--)))))..))....)))))))).))))...)).).))...... ( -23.20, z-score = -0.69, R) >dp4.chr4_group3 9602992 94 - 11692001 GUGGAGUGUGCGCCAGUGUGGCCUUGCC--AGCUAAACUAUUUACCUUAUGUGAUGUUUGUUGUCGCCUUGGCUACAGUUGCCACACAGUCUCUCC---- .....(((((.((.(.(((((((..(((--(((.((((...((((.....)))).))))))))..))...))))))).).))))))).........---- ( -31.00, z-score = -1.94, R) >droPer1.super_1 6717850 94 - 10282868 GUGGAGUGUGCGCCAGUGUGGCCUUGCC--AGCUAAACUAUUUACCUUAUGUGAUGUUUGUUGUCGCCUUGGCUACAGUUGCCACACAGUCUCUCC---- .....(((((.((.(.(((((((..(((--(((.((((...((((.....)))).))))))))..))...))))))).).))))))).........---- ( -31.00, z-score = -1.94, R) >droAna3.scaffold_12943 481322 83 - 5039921 ---UGGUAA-U-CCAGUGUGGCCUUGCC--AACUAAACUAUUUACCUUUUUGGAUGCUUGUUGUUGCCUUGGCUACACUGGAC----GCCCCGC------ ---.(((..-(-(((((((((((..(.(--(((..((.((((((......)))))).))...)))).)..)))))))))))).----)))....------ ( -31.90, z-score = -4.62, R) >consensus GCUGAGUAU_UGCCAGUGUGGCACUGCC__AGCUAAACUAUUUACCUCAUGUGAUGUUUGUUGUCGCCUUGGCUACACUUGUC_CCCGCCCCACC_____ ...((((....((.((((((((...((...(((.((((...((((.....)))).)))))))...))....)))))))).)).....))))......... (-16.35 = -17.49 + 1.14)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:14:57 2011