
| Sequence ID | dm3.chr3LHet |
|---|---|
| Location | 7,803 – 7,865 |
| Length | 62 |
| Max. P | 0.982850 |

| Location | 7,803 – 7,865 |
|---|---|
| Length | 62 |
| Sequences | 3 |
| Columns | 62 |
| Reading direction | forward |
| Mean pairwise identity | 61.83 |
| Shannon entropy | 0.54923 |
| G+C content | 0.54839 |
| Mean single sequence MFE | -16.13 |
| Consensus MFE | -9.24 |
| Energy contribution | -11.13 |
| Covariance contribution | 1.89 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.12 |
| SVM RNA-class probability | 0.982850 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3LHet 7803 62 + 2555491 AUCGUCCGCACUCCAGCACCGGCCAAAUUGCUGACCAAGCAUUUGGCCGGAAGCUCAUACAU .......((......)).((((((((((.(((.....)))))))))))))............ ( -23.40, z-score = -3.93, R) >droSim1.chr3h_random 1333820 54 - 1452968 --------UUCAUUAACACCGGCCCAAUUUCUGACCCAGCCUUUGACCGGAAGCUCUCACAU --------..........((((..(((...(((...)))...))).))))............ ( -8.40, z-score = -1.01, R) >droAna3.scaffold_13280 1158996 62 - 1428709 GCUGACCGCGGUCUACCGCCGUUCAAAUCGCUUACACAGCAUUUGGCCCGAUGCUCGUGCAC ((((((.((((....)))).).)))....))...(((((((((......)))))).)))... ( -16.60, z-score = -0.53, R) >consensus ___G_CCGCACUCUAACACCGGCCAAAUUGCUGACCCAGCAUUUGGCCGGAAGCUCAUACAU ..................((((((((((.(((.....)))))))))))))............ ( -9.24 = -11.13 + 1.89)
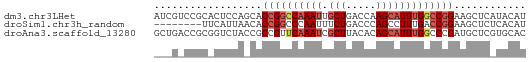
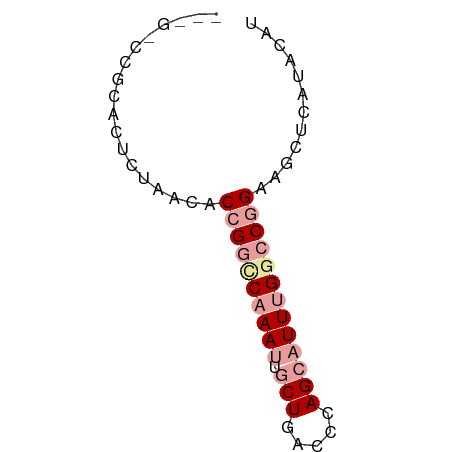
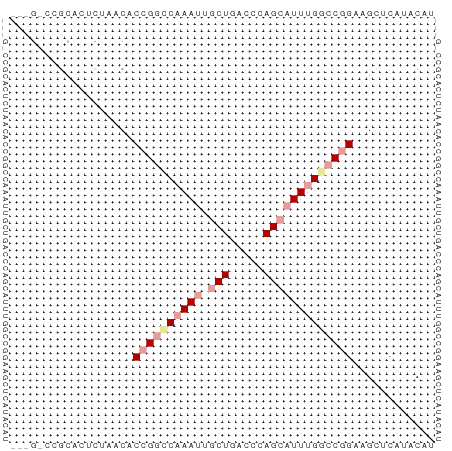
| Location | 7,803 – 7,865 |
|---|---|
| Length | 62 |
| Sequences | 3 |
| Columns | 62 |
| Reading direction | reverse |
| Mean pairwise identity | 61.83 |
| Shannon entropy | 0.54923 |
| G+C content | 0.54839 |
| Mean single sequence MFE | -20.23 |
| Consensus MFE | -10.29 |
| Energy contribution | -12.40 |
| Covariance contribution | 2.11 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.27 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.762605 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
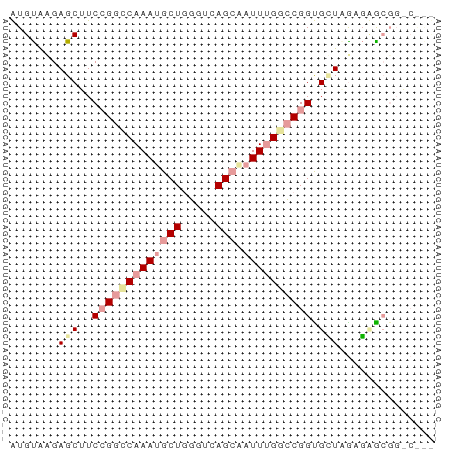
>dm3.chr3LHet 7803 62 - 2555491 AUGUAUGAGCUUCCGGCCAAAUGCUUGGUCAGCAAUUUGGCCGGUGCUGGAGUGCGGACGAU .(((((.(((..(((((((((((((.....))).)))))))))).)))...)))))...... ( -27.20, z-score = -3.07, R) >droSim1.chr3h_random 1333820 54 + 1452968 AUGUGAGAGCUUCCGGUCAAAGGCUGGGUCAGAAAUUGGGCCGGUGUUAAUGAA-------- .......(((..((((((.....(((...)))......)))))).)))......-------- ( -13.10, z-score = -0.32, R) >droAna3.scaffold_13280 1158996 62 + 1428709 GUGCACGAGCAUCGGGCCAAAUGCUGUGUAAGCGAUUUGAACGGCGGUAGACCGCGGUCAGC .(((((.(((((........))))))))))......((((.(.((((....)))).))))). ( -20.40, z-score = -0.43, R) >consensus AUGUAAGAGCUUCCGGCCAAAUGCUGGGUCAGCAAUUUGGCCGGUGCUAGAGAGCGG_C___ .......(((..(((((((((((((.....))).)))))))))).))).............. (-10.29 = -12.40 + 2.11)
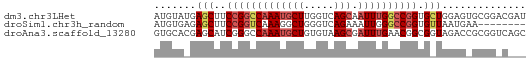

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:49:59 2011