
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 3,909,592 – 3,909,695 |
| Length | 103 |
| Max. P | 0.997583 |

| Location | 3,909,592 – 3,909,695 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 83.89 |
| Shannon entropy | 0.30578 |
| G+C content | 0.50603 |
| Mean single sequence MFE | -35.37 |
| Consensus MFE | -26.67 |
| Energy contribution | -28.02 |
| Covariance contribution | 1.35 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.57 |
| SVM RNA-class probability | 0.950804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
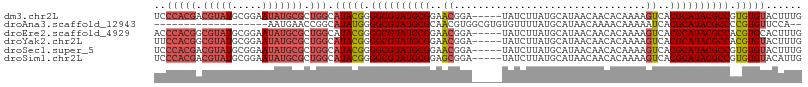
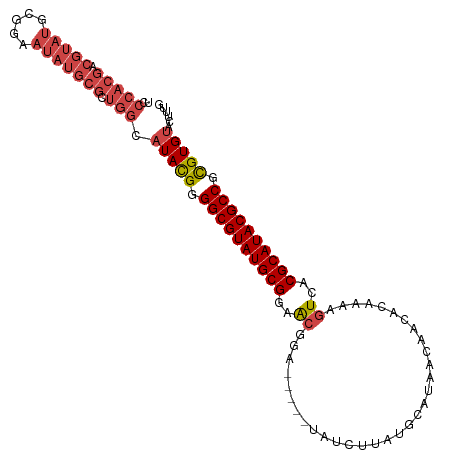
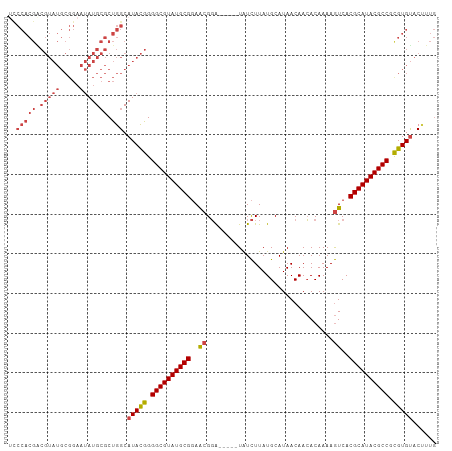
>dm3.chr2L 3909592 103 + 23011544 UCCCACGACGUAUGCGGAAUAUGCGCUGGCAUACGGGGCGUAUGCGGAACGGA-----UAUCUUAUGCAUAACAACACAAAAGUCACGCAUACGCCGUGUGUACUUUG ..(((((.(((((.....))))))).))).(((((.((((((((((..((...-----........................))..)))))))))).)))))...... ( -33.73, z-score = -1.46, R) >droAna3.scaffold_12943 450499 87 - 5039921 -------------------AAUGAACCGGCAUAUGGGGCGUAUGCGCAACGUGGCGUGUGUUUUAUGCAUAACAAAACAAAAAUCACGCAUACGCCCCGUGUUCCA-- -------------------........((.(((((((((((((((((......))(((((((((.(.....).)))))).....)))))))))))))))))).)).-- ( -36.90, z-score = -3.63, R) >droEre2.scaffold_4929 3962059 103 + 26641161 ACCCACGGCGUAUGCGGAAUAUGCGCUGGCAUACGGGGCGUAUGCGGAACGGA-----UAUCUUAUGCAUAACAACACAAAAGUCACGCAUACGCCACGUGCACUUUG .....(((((((((.....)))))))))((((....((((((((((..((...-----........................))..))))))))))..))))...... ( -35.93, z-score = -2.15, R) >droYak2.chr2L 3918237 103 + 22324452 UUCCACGGCGUAUGCGGAAUAUGCGCUGGCAUACGGGGCGUAUGCGGAACGGA-----UAUCUUAUGCAUAACAACACAAAAGUCACGCAUACGCCACGUGUACUUUG .....(((((((((.....)))))))))..(((((.((((((((((..((...-----........................))..)))))))))).)))))...... ( -37.03, z-score = -2.85, R) >droSec1.super_5 2012600 103 + 5866729 UCCCACGACGUAUGCGGAAUAUGCGCUGGCAUACGGGGCGUAUGCGGAACGGA-----UAUCUUAUGCAUAACAACACAAAAGUCACGCAUACGCCGUGUGUACUUUG ..(((((.(((((.....))))))).))).(((((.((((((((((..((...-----........................))..)))))))))).)))))...... ( -33.73, z-score = -1.46, R) >droSim1.chr2L 3864263 103 + 22036055 UCCCACGACGUAUGCGGAAUAUGCGCUGGCAUACGGGGCGUAUGCGGAGCGGA-----UAUCUUAUGCAUAACAACACAAAAGUCACGCAUACGCCGUGUGUACAUUG ...((((.((((((((((((((.((((.(((((((...))))))).).))).)-----)))(((.((..........)).))))).)))))))).))))......... ( -34.90, z-score = -1.30, R) >consensus UCCCACGACGUAUGCGGAAUAUGCGCUGGCAUACGGGGCGUAUGCGGAACGGA_____UAUCUUAUGCAUAACAACACAAAAGUCACGCAUACGCCGCGUGUACUUUG ..(((((.(((((.....))))))).))).(((((.((((((((((..((................................))..)))))))))).)))))...... (-26.67 = -28.02 + 1.35)

| Location | 3,909,592 – 3,909,695 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 83.89 |
| Shannon entropy | 0.30578 |
| G+C content | 0.50603 |
| Mean single sequence MFE | -39.48 |
| Consensus MFE | -29.53 |
| Energy contribution | -30.18 |
| Covariance contribution | 0.65 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.28 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.13 |
| SVM RNA-class probability | 0.997583 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
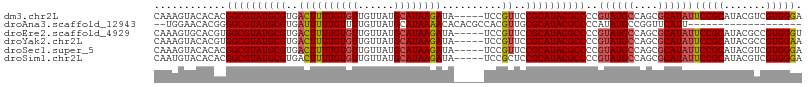
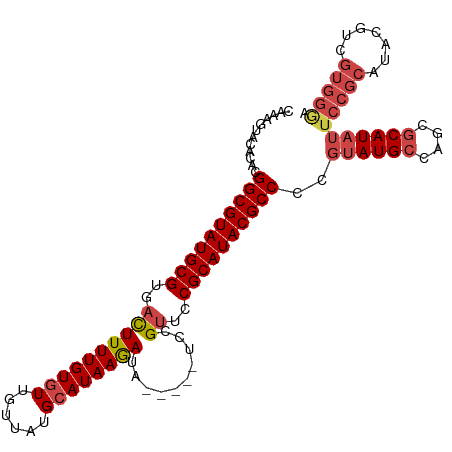
>dm3.chr2L 3909592 103 - 23011544 CAAAGUACACACGGCGUAUGCGUGACUUUUGUGUUGUUAUGCAUAAGAUA-----UCCGUUCCGCAUACGCCCCGUAUGCCAGCGCAUAUUCCGCAUACGUCGUGGGA .........(((((((((((((.((.((((((((......))))))))((-----(.((((..(((((((...))))))).)))).))).)))))))))))))))... ( -41.00, z-score = -3.71, R) >droAna3.scaffold_12943 450499 87 + 5039921 --UGGAACACGGGGCGUAUGCGUGAUUUUUGUUUUGUUAUGCAUAAAACACACGCCACGUUGCGCAUACGCCCCAUAUGCCGGUUCAUU------------------- --.((.....((((((((((((..((...((((((((.....))))))))........))..)))))))))))).....))........------------------- ( -34.50, z-score = -3.17, R) >droEre2.scaffold_4929 3962059 103 - 26641161 CAAAGUGCACGUGGCGUAUGCGUGACUUUUGUGUUGUUAUGCAUAAGAUA-----UCCGUUCCGCAUACGCCCCGUAUGCCAGCGCAUAUUCCGCAUACGCCGUGGGU .........(((((((((((((.((.((((((((......))))))))((-----(.((((..(((((((...))))))).)))).))).)))))))))))))))... ( -38.50, z-score = -2.03, R) >droYak2.chr2L 3918237 103 - 22324452 CAAAGUACACGUGGCGUAUGCGUGACUUUUGUGUUGUUAUGCAUAAGAUA-----UCCGUUCCGCAUACGCCCCGUAUGCCAGCGCAUAUUCCGCAUACGCCGUGGAA .........(((((((((((((.((.((((((((......))))))))((-----(.((((..(((((((...))))))).)))).))).)))))))))))))))... ( -38.50, z-score = -3.05, R) >droSec1.super_5 2012600 103 - 5866729 CAAAGUACACACGGCGUAUGCGUGACUUUUGUGUUGUUAUGCAUAAGAUA-----UCCGUUCCGCAUACGCCCCGUAUGCCAGCGCAUAUUCCGCAUACGUCGUGGGA .........(((((((((((((.((.((((((((......))))))))((-----(.((((..(((((((...))))))).)))).))).)))))))))))))))... ( -41.00, z-score = -3.71, R) >droSim1.chr2L 3864263 103 - 22036055 CAAUGUACACACGGCGUAUGCGUGACUUUUGUGUUGUUAUGCAUAAGAUA-----UCCGCUCCGCAUACGCCCCGUAUGCCAGCGCAUAUUCCGCAUACGUCGUGGGA .........(((((((((((((.((.((((((((......))))))))((-----(.((((..(((((((...))))))).)))).))).)))))))))))))))... ( -43.40, z-score = -3.98, R) >consensus CAAAGUACACACGGCGUAUGCGUGACUUUUGUGUUGUUAUGCAUAAGAUA_____UCCGUUCCGCAUACGCCCCGUAUGCCAGCGCAUAUUCCGCAUACGUCGUGGGA ............((((((((((..((((((((((......))))))))..........))..))))))))))..((((((....))))))(((((.......))))). (-29.53 = -30.18 + 0.65)
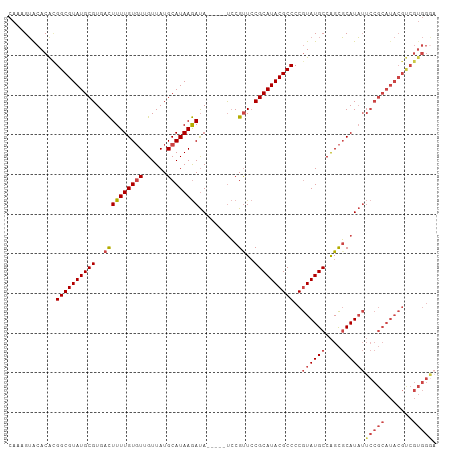
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:14:54 2011