| Sequence ID | dm3.chr2R |
|---|---|
| Location | 20,594,990 – 20,595,082 |
| Length | 92 |
| Max. P | 0.981643 |

| Location | 20,594,990 – 20,595,082 |
|---|---|
| Length | 92 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 67.75 |
| Shannon entropy | 0.64611 |
| G+C content | 0.42608 |
| Mean single sequence MFE | -26.64 |
| Consensus MFE | -13.91 |
| Energy contribution | -14.05 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.47 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.08 |
| SVM RNA-class probability | 0.981643 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
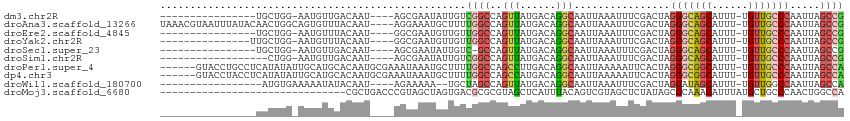
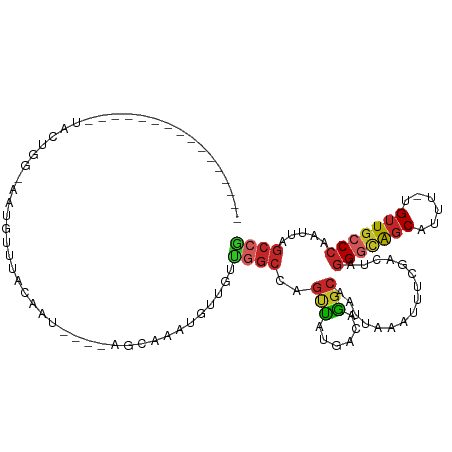
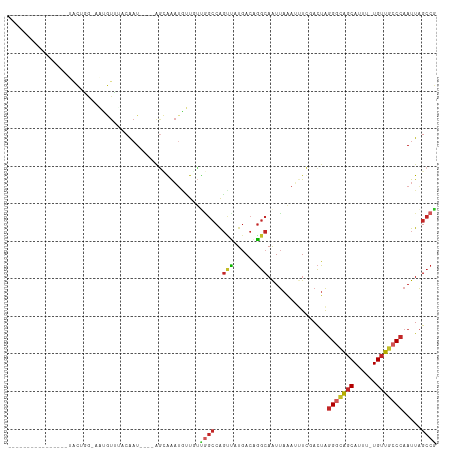
>dm3.chr2R 20594990 92 - 21146708 ----------------UGCUGG-AAUGUUGACAAU----AGCGAAUAUUGUCGGCCAGUUAUGACAGGCAAUUAAAUUUCGACUAGGGCAGCAUUU-UGUUGCCCAAUUAGCCG ----------------.(((((-(((..(((((((----(.....))))))))(((.((....)).)))......))))).....(((((((....-.)))))))....))).. ( -29.60, z-score = -2.72, R) >droAna3.scaffold_13266 7187093 109 - 19884421 UAAACGUAAUUUAUACAACUGGCAGUGUUUACAAU----AGGAAAUGCUUUUGGCCAGUUAUGACAGGCAAUUAAAUUUCGACUAGGGCAGCAUUU-UGUUGCCCAAUUAGCCG .....(((.....)))((((((((((((((.....----...)))))))....)))))))......(((((......))......(((((((....-.))))))).....))). ( -28.80, z-score = -1.88, R) >droEre2.scaffold_4845 21966564 92 - 22589142 ----------------UGCUGG-AAUGUUUACAAU----GGCGAAUGUUGUUGGCCAGUUAUGACAGGCAAUUAAAUUUCGACUAGGGCAGCAUUU-UGUUGCCCAAUUAGCCG ----------------.(((((-..((((.....(----(((.(((...))).)))).....))))..)................(((((((....-.)))))))...)))).. ( -28.10, z-score = -1.68, R) >droYak2.chr2R 20562636 93 - 21139217 ---------------UUGCUGG-AAUGUUUACAAU----GGCGAAUGUUGUUGGCCAGUUAUGACAGGCAAUUAAAUUUCGACUAGGGCAGCAUUU-UGUUGCCCAAUUAGCCG ---------------..(((((-..((((.....(----(((.(((...))).)))).....))))..)................(((((((....-.)))))))...)))).. ( -28.10, z-score = -1.67, R) >droSec1.super_23 461730 91 - 989336 ----------------UGCUGG-AAUGUUGACAAU----AGCGAAUAUUGUC-GCCAGUUAUGACAGGCAAUUAAAUUUCGACUAGGGCAGCAUUU-UGUUGCCCAAUUAGCCG ----------------.(((((-(((...((((((----(.....)))))))-(((.((....)).)))......))))).....(((((((....-.)))))))....))).. ( -29.70, z-score = -3.12, R) >droSim1.chr2R 19050491 90 - 19596830 ------------------CUGG-AAUGUUGACAAU----AGCGAAUAUUGUCGGCCAGUUAUGACAGGCAAUUAAAUUUCGACUAGGGCAGCAUUU-UGUUGCCCAAUUAGCCG ------------------.(((-(((..(((((((----(.....))))))))(((.((....)).)))......))))))....(((((((....-.)))))))......... ( -27.60, z-score = -2.61, R) >droPer1.super_4 931050 107 - 7162766 ------GUACCUGCCUCAUAUAUUGCAUGCACAAUGCGAAAUAAAUGCUUUUGGCCAGCCUUGACAGGCAAUUAAAAAUUCACUAGGGCGGCAUUU-UGUUGCCCAAUUAGCCA ------......((....(((.((((((.....)))))).)))...))...((((..((((....))))................(((((((....-.))))))).....)))) ( -29.60, z-score = -0.88, R) >dp4.chr3 3948337 107 + 19779522 ------GUACCUACCUCAUAUAUUGCAUGCACAAUGCGAAAUAAAUGCUUUUGGCCAGCCAUGACAGGCAAUUAAAAAUUCACUAGGGCGGCAUUU-UGUUGCCCAAUUAGCCA ------............(((.((((((.....)))))).)))........((((..(((......)))................(((((((....-.))))))).....)))) ( -28.50, z-score = -1.07, R) >droWil1.scaffold_180700 2260906 90 + 6630534 -----------------AUGUGAAAAAUAUACAAU----AGAAAAA--UGCUAGCCAGUUAUGACAGGCAAUUAAAUUUCGACUAGGAUAGCAUUU-UGUUGGCCAAUUAGCCA -----------------..................----...((((--((((((((.((....)).)))........(((.....)))))))))))-)..((((......)))) ( -17.60, z-score = -0.92, R) >droMoj3.scaffold_6680 1852411 83 - 24764193 -------------------------------CGCUGACCCGUAGCUAGUGACGCGCGUAGCUCAUUUACAGUCGUAGCUCUAUAGCGCAAACAUUUAUGCUGCCCAACUGGCCA -------------------------------.(((.....(((((.((((..((((((((.....((((....))))..)))).))))...))))...)))))......))).. ( -18.80, z-score = 1.21, R) >consensus ________________UACUGG_AAUGUUUACAAU____AGCAAAUGUUGUUGGCCAGUUAUGACAGGCAAUUAAAUUUCGACUAGGGCAGCAUUU_UGUUGCCCAAUUAGCCG ...................................................((((..(((......)))................((((((((....)))))))).....)))) (-13.91 = -14.05 + 0.14)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:49:01 2011