
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 3,858,053 – 3,858,172 |
| Length | 119 |
| Max. P | 0.985576 |

| Location | 3,858,053 – 3,858,156 |
|---|---|
| Length | 103 |
| Sequences | 11 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 80.19 |
| Shannon entropy | 0.39104 |
| G+C content | 0.50276 |
| Mean single sequence MFE | -32.91 |
| Consensus MFE | -25.13 |
| Energy contribution | -25.13 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.21 |
| SVM RNA-class probability | 0.985576 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
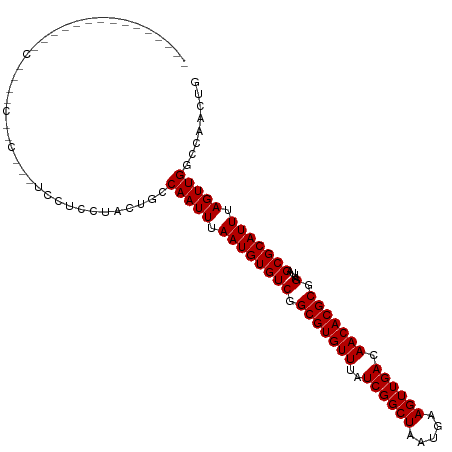
>dm3.chr2L 3858053 103 + 23011544 ------------CCGCGCUGCCGCUGCUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUG ------------..((((....)).))..........(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))))...... ( -34.20, z-score = -1.75, R) >droSim1.chr2L 3812705 103 + 22036055 ------------CCGCGCGGCCGCGGAUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUG ------------(((((....)))))...........(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))))...... ( -39.20, z-score = -2.57, R) >droSec1.super_5 1959639 103 + 5866729 ------------CCGCGCUGCCGCGGAUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUG ------------(((((....)))))...........(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))))...... ( -39.20, z-score = -3.06, R) >droYak2.chr2L 3863637 100 + 22324452 ---------------CCCCACCCUACGUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUG ---------------......................(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))))...... ( -31.50, z-score = -2.95, R) >droEre2.scaffold_4929 3909633 92 + 26641161 -----------------------CGGGUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUG -----------------------.(((....)))...(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))))...... ( -33.30, z-score = -3.16, R) >dp4.chr4_group3 6835496 115 - 11692001 CGCCAAACUGCUGAGCGUCUCGGCCCUGCAGCCUGCUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUG .((....((((.(.((......)).).))))...)).(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))))...... ( -40.20, z-score = -1.39, R) >droPer1.super_1 3935296 115 - 10282868 CGCCAAACUGCUGAGCGUCUCGGCCCUGCAGCCUGCUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUG .((....((((.(.((......)).).))))...)).(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))))...... ( -40.20, z-score = -1.39, R) >droWil1.scaffold_180708 9755255 82 + 12563649 --------------------------------AGUCCGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUG-CCAACUG --------------------------------(((..((.((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))-)..))). ( -26.30, z-score = -2.19, R) >droVir3.scaffold_13246 2437254 85 + 2672878 -----------------------------GAGGAGCACCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUG-CCAACUG -----------------------------..((......(((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))-))..... ( -24.80, z-score = -1.46, R) >droMoj3.scaffold_6500 26517683 83 + 32352404 -------------------------------GGGGCACCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUG-CCAACUG -------------------------------..((((....(((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))-))..... ( -26.90, z-score = -1.91, R) >droGri2.scaffold_15252 5887568 83 + 17193109 -------------------------------GGGGCCACCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUG-CCAACUG -------------------------------..(((....((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))-))..... ( -26.20, z-score = -1.67, R) >consensus _______________C____C__C___UCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUG .......................................(((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))........ (-25.13 = -25.13 + -0.00)


| Location | 3,858,058 – 3,858,172 |
|---|---|
| Length | 114 |
| Sequences | 12 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.69 |
| Shannon entropy | 0.39957 |
| G+C content | 0.51340 |
| Mean single sequence MFE | -37.61 |
| Consensus MFE | -25.17 |
| Energy contribution | -25.17 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.48 |
| SVM RNA-class probability | 0.941893 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
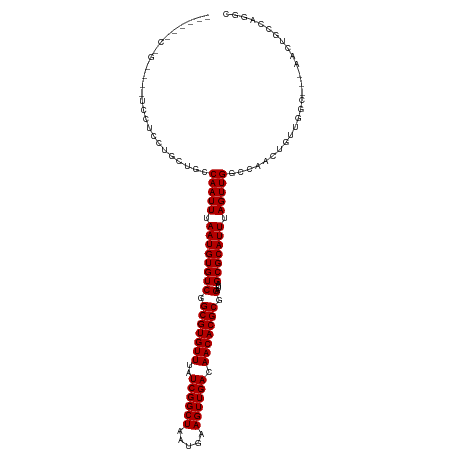
>dm3.chr2L 3858058 114 + 23011544 ---CUGCCGCUGCUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUGUUGGC---AACUGCCAGGC ---..(((...............(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).))))))).......((((---....))))))) ( -41.00, z-score = -2.13, R) >droSim1.chr2L 3812710 114 + 22036055 ---CGGCCGCGGAUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUGUUGGC---AACUGCCAGGC ---..(((..((.....))....(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).))))))).......((((---....))))))) ( -41.90, z-score = -2.00, R) >droSec1.super_5 1959644 114 + 5866729 ---CUGCCGCGGAUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUGUUGGC---AACUGCCAGGC ---..(((..((.....))....(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).))))))).......((((---....))))))) ( -41.90, z-score = -2.33, R) >droYak2.chr2L 3863639 114 + 22324452 ---CCACCCUACGUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUGUUGGC---AACUGCCAGGC ---....(((.............(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))))........(((---....)))))). ( -38.10, z-score = -2.08, R) >droEre2.scaffold_4929 3909634 107 + 26641161 ----------GGGUCCUCCUACUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUGUUGGC---AACUGGCAGGC ----------(((....)))...(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))))...((((..(.---..)..)))).. ( -39.30, z-score = -2.44, R) >droAna3.scaffold_12916 1862859 117 + 16180835 CCCCCGCCGGCUCUCCCUGCGCUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUGUUGGC---AACUGCCGAGG ...(((((((((.....(((((((............(((.(((((((..((((((.....)))))).))))))).))))))))))...)))))))......(((((---....))))))) ( -43.10, z-score = -1.90, R) >dp4.chr4_group3 6835513 114 - 11692001 ---UCUCGGCCCUGCAGCCUGCUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUGUUGGC---CACUGACGGAG ---....((((..((((..((..(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).))))))))).)))).)))---).......... ( -44.10, z-score = -2.62, R) >droPer1.super_1 3935313 114 - 10282868 ---UCUCGGCCCUGCAGCCUGCUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUGUUGGC---CACUGACGGAG ---....((((..((((..((..(((((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).))))))))).)))).)))---).......... ( -44.10, z-score = -2.62, R) >droWil1.scaffold_180708 9755257 90 + 12563649 --------------------UCCGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUG-CCAACUGGCCUC---GACUU------ --------------------...((((.......(((((.(((((((..((((((.....)))))).))))))).)))..))(((......))-)....))))...---.....------ ( -26.60, z-score = -1.19, R) >droVir3.scaffold_13246 2437258 95 + 2672878 -------------------AGCACCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUG-CCAACUG-CUGC---GACUCCAGGC- -------------------((((..(((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))-.....))-))..---..........- ( -27.40, z-score = -0.49, R) >droMoj3.scaffold_6500 26517685 95 + 32352404 -------------------GGCACCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUG-CCAACUG-CUGC---GACUCCAGGC- -------------------(((((..........))))).(((((((..((((((.....)))))).)))))))(((....((((..((((..-...))))-.)))---)..)))....- ( -28.30, z-score = -0.61, R) >droGri2.scaffold_15252 5887570 100 + 17193109 -------------------GGCCACCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUG-CCAACUGCUGGCUCUGACUGCCAGUC -------------------(((....((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))-))....((((((.......)))))). ( -35.50, z-score = -2.40, R) >consensus ______C_G____UCCUCCUGCUGCCAAUUUAAUGUGUCGGCGUGUUUAUCGGCUAAUGAAGUUGACAACACGCGGAUUAGCGCAUUUAGUUGGCCAACUGUUGGC___AACUGCCAGGC .........................(((((.((((((((.(((((((..((((((.....)))))).))))))).)....))))))).)))))........................... (-25.17 = -25.17 + 0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:14:49 2011