
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 20,296,855 – 20,296,954 |
| Length | 99 |
| Max. P | 0.897856 |

| Location | 20,296,855 – 20,296,954 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 73.77 |
| Shannon entropy | 0.42823 |
| G+C content | 0.48314 |
| Mean single sequence MFE | -29.95 |
| Consensus MFE | -20.61 |
| Energy contribution | -21.05 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.897856 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
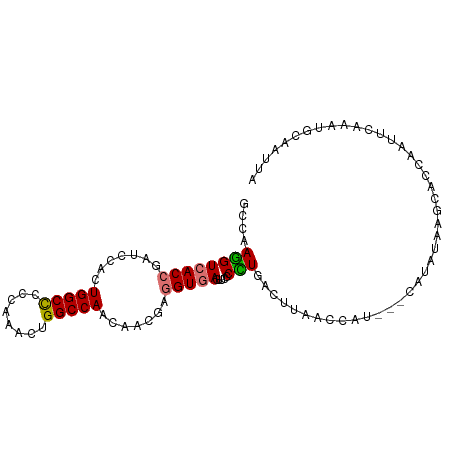
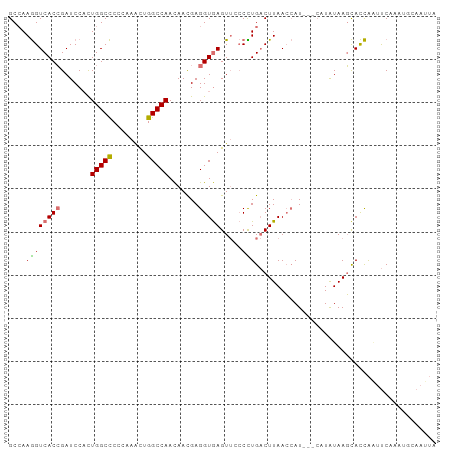
>dm3.chr2R 20296855 99 + 21146708 UAAUUCCAUAUGAACUGGUGCUUAUAUGCUAAUGGUUAAGUCAAGGUAACUCACCUCGUUGUUGGCCAGCUUGGGGGCCAGUGGAUCGGUGACCUUGGC ............(((((..((......))...)))))..(((((((....(((((...(..((((((........))))))..)...)))))))))))) ( -32.20, z-score = -1.24, R) >droSim1.chr2R 18750168 94 + 19596830 UAAUUGGAUUUGAAUUGGUGCUUAUA-----AUGGUUAAGUCAGGGGAACUCACCUCGUUGUUGGCCAGUUUGGGGGCCAGUGGAUCGGUGACCUUGGC ......((((...((((((.((((..-----.(((((((....((((......))))....)))))))...)))).)))))).))))(....)...... ( -28.30, z-score = -0.88, R) >droSec1.super_23 167089 99 + 989336 UAAUUGCAUUUGAAUUGGUGCUUAUAUGCUAAUGGUUAAGUCAGGGGAACUCACCUCGUUGUUGGCCAGCUUGGGGGCCAGUGGAUCGGUGACCUUGGC .....(((((......)))))..................(((((((....(((((...(..((((((........))))))..)...)))))))))))) ( -31.60, z-score = -0.90, R) >droAna3.scaffold_13266 15834312 82 + 19884421 -----------GGGGCAAUAGUUACAGG---AUUCUUAAGGCAAG---ACUUACCUCAUUAUUGGCCAAUCGUGGAGCCAAAGGAUCCGUGACUUUUGC -----------...((((.((((((.((---((((((..(((...---.....((........))(((....))).))).)))))))))))))).)))) ( -27.70, z-score = -3.00, R) >consensus UAAUUGCAUUUGAAUUGGUGCUUAUAUG___AUGGUUAAGUCAAGGGAACUCACCUCGUUGUUGGCCAGCUUGGGGGCCAGUGGAUCGGUGACCUUGGC .......................................(((((((....(((((...(..((((((........))))))..)...)))))))))))) (-20.61 = -21.05 + 0.44)



| Location | 20,296,855 – 20,296,954 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 73.77 |
| Shannon entropy | 0.42823 |
| G+C content | 0.48314 |
| Mean single sequence MFE | -21.66 |
| Consensus MFE | -12.59 |
| Energy contribution | -12.96 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.551419 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 20296855 99 - 21146708 GCCAAGGUCACCGAUCCACUGGCCCCCAAGCUGGCCAACAACGAGGUGAGUUACCUUGACUUAACCAUUAGCAUAUAAGCACCAGUUCAUAUGGAAUUA ..((((((((((.......(((((........))))).......)))))....))))).............(((((.(((....))).)))))...... ( -24.94, z-score = -1.33, R) >droSim1.chr2R 18750168 94 - 19596830 GCCAAGGUCACCGAUCCACUGGCCCCCAAACUGGCCAACAACGAGGUGAGUUCCCCUGACUUAACCAU-----UAUAAGCACCAAUUCAAAUCCAAUUA .....(((((((.......(((((........))))).......))))).......((.((((.....-----..))))))))................ ( -17.54, z-score = -0.85, R) >droSec1.super_23 167089 99 - 989336 GCCAAGGUCACCGAUCCACUGGCCCCCAAGCUGGCCAACAACGAGGUGAGUUCCCCUGACUUAACCAUUAGCAUAUAAGCACCAAUUCAAAUGCAAUUA ((((.(((((((.......(((((........))))).......)))))....)).))............((......))............))..... ( -19.14, z-score = -0.08, R) >droAna3.scaffold_13266 15834312 82 - 19884421 GCAAAAGUCACGGAUCCUUUGGCUCCACGAUUGGCCAAUAAUGAGGUAAGU---CUUGCCUUAAGAAU---CCUGUAACUAUUGCCCC----------- ((((.(((.((((((.((((((((........))))))...(((((((...---..))))))))).))---)).)).))).))))...----------- ( -25.00, z-score = -3.19, R) >consensus GCCAAGGUCACCGAUCCACUGGCCCCCAAACUGGCCAACAACGAGGUGAGUUCCCCUGACUUAACCAU___CAUAUAAGCACCAAUUCAAAUGCAAUUA ....((.(((((.......(((((........))))).......))))).))............................................... (-12.59 = -12.96 + 0.37)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:48:24 2011