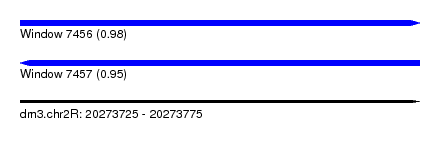

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 20,273,725 – 20,273,775 |
| Length | 50 |
| Max. P | 0.981846 |

| Location | 20,273,725 – 20,273,775 |
|---|---|
| Length | 50 |
| Sequences | 10 |
| Columns | 51 |
| Reading direction | forward |
| Mean pairwise identity | 95.35 |
| Shannon entropy | 0.10180 |
| G+C content | 0.40325 |
| Mean single sequence MFE | -11.09 |
| Consensus MFE | -10.53 |
| Energy contribution | -10.31 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.09 |
| SVM RNA-class probability | 0.981846 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
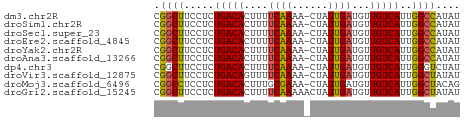
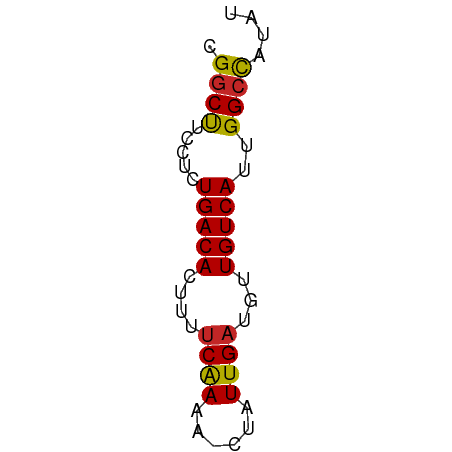
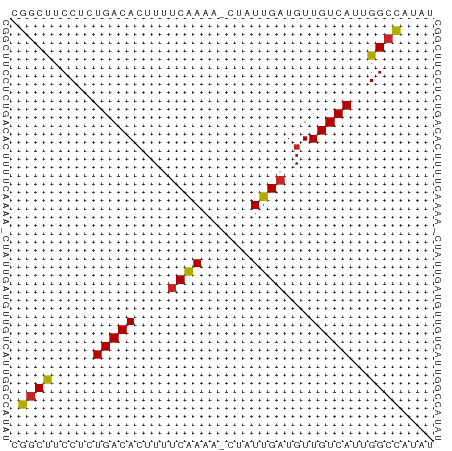
>dm3.chr2R 20273725 50 + 21146708 CGGCUUCCUCUGACACUUUUCAAAA-CUAUUGAUGUUGUCAUUGGCCAUAU .((((.....(((((....((((..-...))))...)))))..)))).... ( -11.90, z-score = -2.78, R) >droSim1.chr2R 18724921 50 + 19596830 CGGCUUCCUCUGACACUUUUCAAAA-CUAUUGAUGUUGUCAUUGGCCAUAU .((((.....(((((....((((..-...))))...)))))..)))).... ( -11.90, z-score = -2.78, R) >droSec1.super_23 144177 50 + 989336 CGGCUUCCUCUGACACUUUUCAAAA-CUAUUGAUGUUGUCAUUGGCCAUAU .((((.....(((((....((((..-...))))...)))))..)))).... ( -11.90, z-score = -2.78, R) >droEre2.scaffold_4845 21631974 50 + 22589142 CGGCUUCCUCUGACACUUUUCAAAA-CUAUUGAUGUUGUCAUUGGCCAUAU .((((.....(((((....((((..-...))))...)))))..)))).... ( -11.90, z-score = -2.78, R) >droYak2.chr2R 20234747 50 + 21139217 CGGCUUCCUCUGACACUUUUCAAAA-CUAUUGAUGUUGUCAUUGGCCAUAU .((((.....(((((....((((..-...))))...)))))..)))).... ( -11.90, z-score = -2.78, R) >droAna3.scaffold_13266 5737169 50 - 19884421 CGGCUUCCUCUGACACUUUUCAAAA-CUAUUGAUGUUGUCAUUGGCCAUAU .((((.....(((((....((((..-...))))...)))))..)))).... ( -11.90, z-score = -2.78, R) >dp4.chr3 1378201 50 - 19779522 CGGCUUCCUCUGACACUUUUCAAAA-CUAUUGAUGUUGUCAUUGGGUCUAU .(((..((..(((((....((((..-...))))...)))))..)))))... ( -10.60, z-score = -1.98, R) >droVir3.scaffold_12875 7389301 50 + 20611582 CGGCUUCCUCUGACAGUUUUCAAAA-CUAUUGAUGUUGUCAUUGGCUAUAU .((((.....((((((...((((..-...))))..))))))..)))).... ( -10.20, z-score = -1.69, R) >droMoj3.scaffold_6496 26185151 50 + 26866924 CGGCCUCCUCUGACACUUUGCGAAA-CUAUUGAUGUUGUCAUUGGCUACAG .((((.....(((((.....(((..-...)))....)))))..)))).... ( -9.30, z-score = -0.82, R) >droGri2.scaffold_15245 938631 51 + 18325388 CGGCUUCCUCUGACACUUUUCAAAAACUAUUGAUGUUGUCAUUGGCUAUAU .((((.....(((((....((((......))))...)))))..)))).... ( -9.40, z-score = -2.06, R) >consensus CGGCUUCCUCUGACACUUUUCAAAA_CUAUUGAUGUUGUCAUUGGCCAUAU .((((.....(((((....((((......))))...)))))..)))).... (-10.53 = -10.31 + -0.22)
| Location | 20,273,725 – 20,273,775 |
|---|---|
| Length | 50 |
| Sequences | 10 |
| Columns | 51 |
| Reading direction | reverse |
| Mean pairwise identity | 95.35 |
| Shannon entropy | 0.10180 |
| G+C content | 0.40325 |
| Mean single sequence MFE | -10.82 |
| Consensus MFE | -9.84 |
| Energy contribution | -10.04 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.946084 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
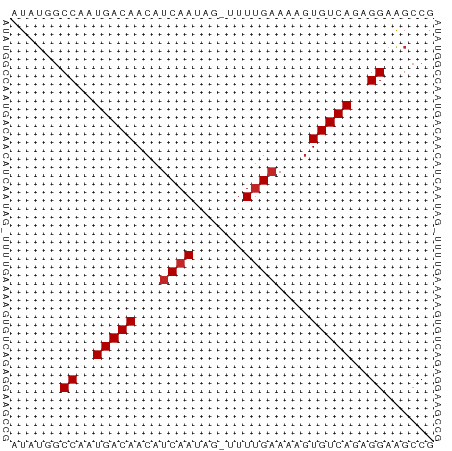
>dm3.chr2R 20273725 50 - 21146708 AUAUGGCCAAUGACAACAUCAAUAG-UUUUGAAAAGUGUCAGAGGAAGCCG ....(((...(((((...((((...-..))))....)))))......))). ( -11.60, z-score = -2.32, R) >droSim1.chr2R 18724921 50 - 19596830 AUAUGGCCAAUGACAACAUCAAUAG-UUUUGAAAAGUGUCAGAGGAAGCCG ....(((...(((((...((((...-..))))....)))))......))). ( -11.60, z-score = -2.32, R) >droSec1.super_23 144177 50 - 989336 AUAUGGCCAAUGACAACAUCAAUAG-UUUUGAAAAGUGUCAGAGGAAGCCG ....(((...(((((...((((...-..))))....)))))......))). ( -11.60, z-score = -2.32, R) >droEre2.scaffold_4845 21631974 50 - 22589142 AUAUGGCCAAUGACAACAUCAAUAG-UUUUGAAAAGUGUCAGAGGAAGCCG ....(((...(((((...((((...-..))))....)))))......))). ( -11.60, z-score = -2.32, R) >droYak2.chr2R 20234747 50 - 21139217 AUAUGGCCAAUGACAACAUCAAUAG-UUUUGAAAAGUGUCAGAGGAAGCCG ....(((...(((((...((((...-..))))....)))))......))). ( -11.60, z-score = -2.32, R) >droAna3.scaffold_13266 5737169 50 + 19884421 AUAUGGCCAAUGACAACAUCAAUAG-UUUUGAAAAGUGUCAGAGGAAGCCG ....(((...(((((...((((...-..))))....)))))......))). ( -11.60, z-score = -2.32, R) >dp4.chr3 1378201 50 + 19779522 AUAGACCCAAUGACAACAUCAAUAG-UUUUGAAAAGUGUCAGAGGAAGCCG ......((..(((((...((((...-..))))....)))))..))...... ( -10.60, z-score = -2.87, R) >droVir3.scaffold_12875 7389301 50 - 20611582 AUAUAGCCAAUGACAACAUCAAUAG-UUUUGAAAACUGUCAGAGGAAGCCG ......((..(((((...((((...-..))))....)))))..))...... ( -9.40, z-score = -2.04, R) >droMoj3.scaffold_6496 26185151 50 - 26866924 CUGUAGCCAAUGACAACAUCAAUAG-UUUCGCAAAGUGUCAGAGGAGGCCG ..((..((..(((((..........-..........)))))..))..)).. ( -7.95, z-score = 0.02, R) >droGri2.scaffold_15245 938631 51 - 18325388 AUAUAGCCAAUGACAACAUCAAUAGUUUUUGAAAAGUGUCAGAGGAAGCCG ......((..(((((...((((......))))....)))))..))...... ( -10.70, z-score = -2.73, R) >consensus AUAUGGCCAAUGACAACAUCAAUAG_UUUUGAAAAGUGUCAGAGGAAGCCG ......((..(((((...((((......))))....)))))..))...... ( -9.84 = -10.04 + 0.20)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:48:15 2011