
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 20,269,653 – 20,269,746 |
| Length | 93 |
| Max. P | 0.814781 |

| Location | 20,269,653 – 20,269,746 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 91.23 |
| Shannon entropy | 0.15088 |
| G+C content | 0.38568 |
| Mean single sequence MFE | -22.74 |
| Consensus MFE | -17.02 |
| Energy contribution | -17.02 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.814781 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
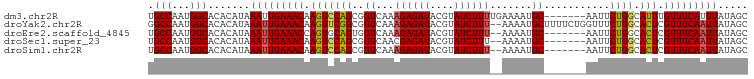
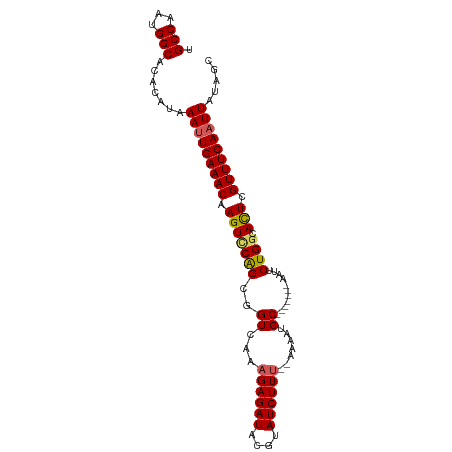
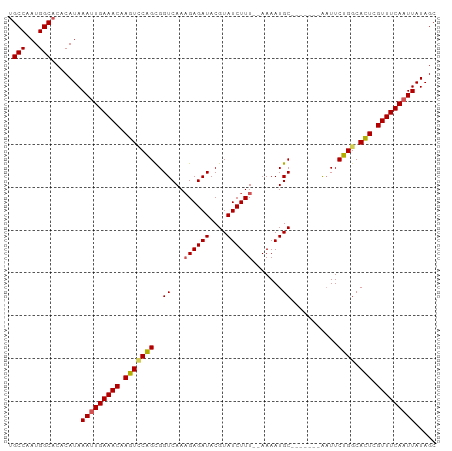
>dm3.chr2R 20269653 93 + 21146708 UGCCAAUGGCACACAUAAAUUGAAACAAGUCCAGCGGUCAAAGAGAUACGUAUCUUUUGAAAAUGC-------AAUUCUGGCAUUUGUUUCAUUUAUAGC ((((...))))...((((((.(((((((((((((((.((((.(((((....)))))))))...)))-------.....))).)))))))))))))))... ( -25.00, z-score = -2.89, R) >droYak2.chr2R 20230680 98 + 21139217 GGCCAAUGGCACACAUAAAUUGAAACAAGUUCGGCGGUCAAAGAGAUACGUAUCUUU--AAAAUGCUUUUCUGGUUUCUGGCACUCGUUUCAAUUAUAGC .(((...))).......(((((((((.(((((((.(..(((((((.....((....)--).....))))).))..).)))).))).)))))))))..... ( -21.20, z-score = -1.21, R) >droEre2.scaffold_4845 21627899 91 + 22589142 UGCCAAUGGCACACAUAAAUUGAAACCAGUGCAGUGGUCAAAGAGAUACGUAUCUUU--AAAAUGC-------AAUUCUGGCACUCGUUUCAAUUAUAGC ((((...))))......(((((((((.(((((((..((...((((((....))))))--.....))-------....)).))))).)))))))))..... ( -23.10, z-score = -2.25, R) >droSec1.super_23 140177 91 + 989336 UGCCAAUGGCACACAUAAAUUGAAACAAGUCCAGCGGUCAACGAGAUACGUAUCUUU--AAAAUGC-------AAUUCUGGCACUCGUUUCAAUUAUAGC ((((...))))......(((((((((.(((((((..((....(((((....))))).--.....))-------....)))).))).)))))))))..... ( -21.80, z-score = -2.45, R) >droSim1.chr2R 18720920 91 + 19596830 UGCCAAUGGCACACAUAAAUUGAAACAAGUCCAGCGGUCAAAGAGAUACGUAUCUUU--AAAAUGC-------AAUUCUGGCACUCGUUUCAAUUAUAGC ((((...))))......(((((((((.(((((((..((...((((((....))))))--.....))-------....)))).))).)))))))))..... ( -22.60, z-score = -2.88, R) >consensus UGCCAAUGGCACACAUAAAUUGAAACAAGUCCAGCGGUCAAAGAGAUACGUAUCUUU__AAAAUGC_______AAUUCUGGCACUCGUUUCAAUUAUAGC .(((...))).......(((((((((.(((((((((.....((((((....))))))......)))............))).))).)))))))))..... (-17.02 = -17.02 + -0.00)
| Location | 20,269,653 – 20,269,746 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 91.23 |
| Shannon entropy | 0.15088 |
| G+C content | 0.38568 |
| Mean single sequence MFE | -23.85 |
| Consensus MFE | -17.15 |
| Energy contribution | -18.43 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.690109 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
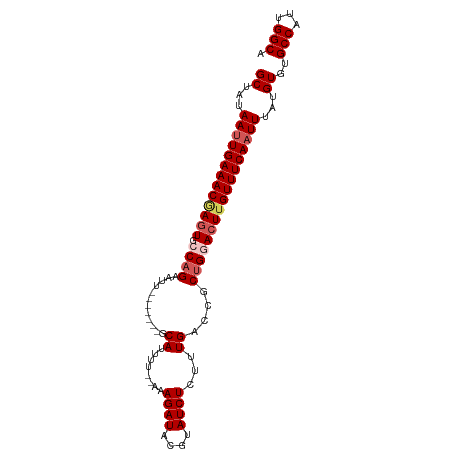
>dm3.chr2R 20269653 93 - 21146708 GCUAUAAAUGAAACAAAUGCCAGAAUU-------GCAUUUUCAAAAGAUACGUAUCUCUUUGACCGCUGGACUUGUUUCAAUUUAUGUGUGCCAUUGGCA ((((((((((((((((...((((....-------......(((((((((....)))).)))))...))))..))))))).))))))).))(((...))). ( -21.82, z-score = -1.77, R) >droYak2.chr2R 20230680 98 - 21139217 GCUAUAAUUGAAACGAGUGCCAGAAACCAGAAAAGCAUUUU--AAAGAUACGUAUCUCUUUGACCGCCGAACUUGUUUCAAUUUAUGUGUGCCAUUGGCC ((...(((((((((((((((..............))...((--(((((........))))))).......)))))))))))))...))..(((...))). ( -20.24, z-score = -1.47, R) >droEre2.scaffold_4845 21627899 91 - 22589142 GCUAUAAUUGAAACGAGUGCCAGAAUU-------GCAUUUU--AAAGAUACGUAUCUCUUUGACCACUGCACUGGUUUCAAUUUAUGUGUGCCAUUGGCA ((...(((((((((.(((((.((...(-------(....((--(((((........))))))).))))))))).)))))))))...)).((((...)))) ( -24.50, z-score = -2.35, R) >droSec1.super_23 140177 91 - 989336 GCUAUAAUUGAAACGAGUGCCAGAAUU-------GCAUUUU--AAAGAUACGUAUCUCGUUGACCGCUGGACUUGUUUCAAUUUAUGUGUGCCAUUGGCA ((...(((((((((((((.((((....-------.((....--..((((....))))...))....)))))))))))))))))...)).((((...)))) ( -26.00, z-score = -2.57, R) >droSim1.chr2R 18720920 91 - 19596830 GCUAUAAUUGAAACGAGUGCCAGAAUU-------GCAUUUU--AAAGAUACGUAUCUCUUUGACCGCUGGACUUGUUUCAAUUUAUGUGUGCCAUUGGCA ((...(((((((((((((.((((....-------.....((--(((((........)))))))...)))))))))))))))))...)).((((...)))) ( -26.70, z-score = -3.25, R) >consensus GCUAUAAUUGAAACGAGUGCCAGAAUU_______GCAUUUU__AAAGAUACGUAUCUCUUUGACCGCUGGACUUGUUUCAAUUUAUGUGUGCCAUUGGCA ((...(((((((((((((.((((....................(((((........))))).....)))))))))))))))))...))..(((...))). (-17.15 = -18.43 + 1.28)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:48:14 2011