
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 20,102,892 – 20,103,006 |
| Length | 114 |
| Max. P | 0.500000 |

| Location | 20,102,892 – 20,103,006 |
|---|---|
| Length | 114 |
| Sequences | 9 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.64 |
| Shannon entropy | 0.45885 |
| G+C content | 0.67073 |
| Mean single sequence MFE | -51.67 |
| Consensus MFE | -23.84 |
| Energy contribution | -23.20 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.63 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
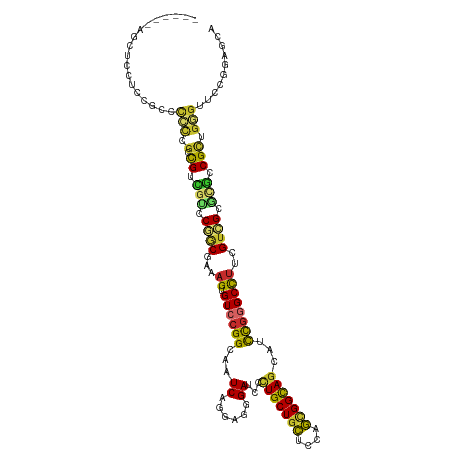
>dm3.chr2R 20102892 114 + 21146708 ------AGCUCCUCCGCCCCCCGCCGUCGUCCGGCGAAAGUGUCCGGCAAUCAGGAGGGAUCCCUGCUGCUCCAGCGGCAGCAUCCGGGCUUUCGUCGCGCGCCGUUGGGUUCCGGAGCA ------.(((((.......((((.((.(((.((((((((((..((((......(((....)))(((((((....)))))))...)))))))))))))).))).)).))))....))))). ( -54.40, z-score = -1.68, R) >droMoj3.scaffold_6496 25906851 120 + 26866924 UGCAGCACCGCCUGUCGAUGUUAUUGACGGCCUGCGAAAGUGUCCGGCAAUCAGCAGAGAGCCCUGCUGGUUCAUGGGCAGCAUCCUGGCCUUGGUCGCCUGCCGCUGCGUGCCAGCGCA .(((((...(((.((((((...))))))((.(.((....))).)))))(((((((((......)))))))))....((((((..((.......))..).))))))))))(((....))). ( -49.70, z-score = -0.03, R) >droWil1.scaffold_180701 3640716 103 - 3904529 --------------CAGAUGCUGUUGU---CCUUCGAAAGUGUCCGGCAAUCAGUAACGAGCCCUGCUGCUCCAGCGGCACCAUUCGUGCCCUUGUGGCCUGACGUUGCGUGCCAGAGCA --------------..((((((.(((.---....))).)))))).((((....(((((((((...)))((..(((.(((((.....))))).)))..))....)))))).))))...... ( -35.10, z-score = -0.75, R) >droPer1.super_34 398989 103 - 916997 -----------------CCACUGGCGUGCUCCUGCCAAAGUGUCCGGCAAUCAGCAGGGAACCCUGCUGCUCCAGUGGCAGCAUUUGUGCCUUUGUCGCCCUACGCUGGGUGCCGGAGAA -----------------.((((((((......))))..))))(((((((..(((((((....))))))).(((((((((.(((..........))).)))....)))))))))))))... ( -45.60, z-score = -1.90, R) >droAna3.scaffold_13266 5581395 110 - 19884421 ----------CAUCCACCACCGGGCGAUGUCCUGCGAAAGUGUCCGGCAAUCAGGAGCGAGCCUUGCUGCUCGAGCGGUAGCAUUCGGGCCUUGGUGGCACGUCGCUGGGUCCCGGAGCA ----------..(((...(((.((((((((.(..(..(((.((((((.......((((.(((...)))))))..((....))..))))))))).)..).)))))))).)))...)))... ( -46.00, z-score = -0.07, R) >droEre2.scaffold_4845 21465658 114 + 22589142 ------AGCUCCGCCGCCCCCCGCCGUCGUCCGACGAAAGUGUCCGGCAAUCAGGAGGGAUCCCUGCUGCUCCAGCGGCAGCAUCCGGGCUUUCGUCGCGCGCCGCUGGGUUCCGGAGCA ------.((((((..((((...((.(.(((.((((((((((..((((......(((....)))(((((((....)))))))...)))))))))))))).)))).)).))))..)))))). ( -59.70, z-score = -2.93, R) >droYak2.chr2R 20067297 114 + 21139217 ------AGCUCCUCCACCCGGCGCCGUCGUCCGGCGAAAGUGUCCGGCAAUCAGCAGGGAUCCCUGCUGCUCCAGUGGCAGCAUCCGGGCCUUCGUCGCGCGCCGCUGGGUUCCGGAGCA ------.(((((...((((((((.((.((..(((((((.(.((((((.....((((((....))))))(((((...)).)))..)))))))))))))))))).))))))))...))))). ( -62.70, z-score = -3.94, R) >droSec1.super_5 342218 114 + 5866729 ------AGCUCCUCCGCCCCCCGCCGUCGUCCGGCGAAAGUGUCCGGCAAUCAGGAGGGAUCCCUGCUGCUCCAGCGGCAGCAUCCGGGCUUUCGUCGCGCGCCGCUGGGUUCCGGAGCA ------.(((((...((((...((.(.(((.((((((((((..((((......(((....)))(((((((....)))))))...)))))))))))))).)))).)).))))...))))). ( -55.90, z-score = -1.77, R) >droSim1.chr2R 18584027 114 + 19596830 ------AGCUCCUCCGCCCCCCGCCGUCGUCCGGCGAAAGUGUCCGGCAAUCAGGAGGGAUCCCUGCUGCUCCAGCGGCAGCAUCCGGGCUUUCGUCGCGCGCCGCUGGGUUCCGGAGCA ------.(((((...((((...((.(.(((.((((((((((..((((......(((....)))(((((((....)))))))...)))))))))))))).)))).)).))))...))))). ( -55.90, z-score = -1.77, R) >consensus ______AGCUCCUCCGCCCCCCGCCGUCGUCCGGCGAAAGUGUCCGGCAAUCAGGAGGGAUCCCUGCUGCUCCAGCGGCAGCAUCCGGGCCUUCGUCGCGCGCCGCUGGGUUCCGGAGCA ..................(((.(.((.(((.((((...((.((((((...((......))...(((((((....)))))))...))))))))..)))).))).))).))).......... (-23.84 = -23.20 + -0.64)
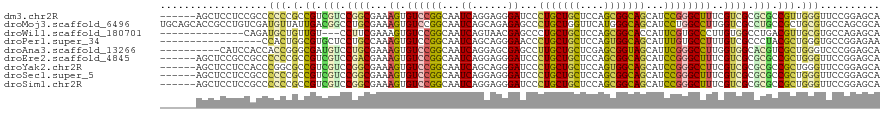


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:47:38 2011