
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 19,921,365 – 19,921,454 |
| Length | 89 |
| Max. P | 0.937055 |

| Location | 19,921,365 – 19,921,454 |
|---|---|
| Length | 89 |
| Sequences | 9 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 88.03 |
| Shannon entropy | 0.25005 |
| G+C content | 0.33749 |
| Mean single sequence MFE | -20.52 |
| Consensus MFE | -17.44 |
| Energy contribution | -17.22 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.41 |
| SVM RNA-class probability | 0.937055 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
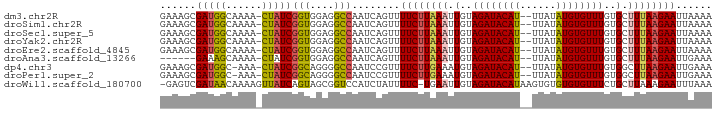
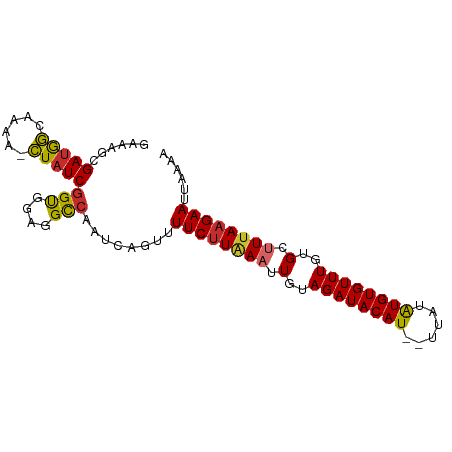
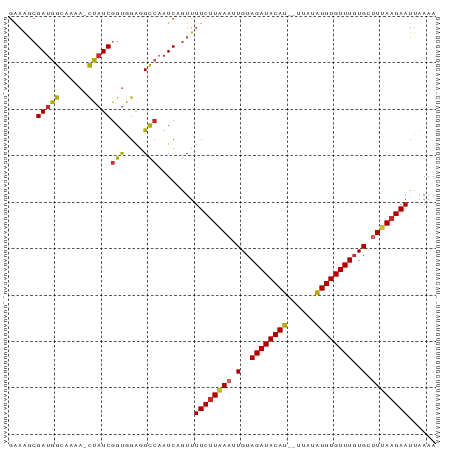
>dm3.chr2R 19921365 89 - 21146708 GAAAGCGAUGGCAAAA-CUAUCGGUGGAGGCCAAUCAGUUUUCUUAAAUUGUAGAUACAU--UUAUAUGUGUUUGUGCUUUAAGAAUUAAAA ......((((((...(-(.....))....))).)))....((((((((.(..((((((((--....))))))))..).))))))))...... ( -20.90, z-score = -2.17, R) >droSim1.chr2R 18458028 89 - 19596830 GAAAGCGAUGGCAAAA-CUAUCGGUGGAGGCCAAUCAGUUUUCUUAAAUUGUAGAUACAU--UUAUAUGUGUUUGUGCUUUAAGAAUUAAAA ......((((((...(-(.....))....))).)))....((((((((.(..((((((((--....))))))))..).))))))))...... ( -20.90, z-score = -2.17, R) >droSec1.super_5 174527 89 - 5866729 GAAAGCGAUGGCAAAA-CUAUCGGUGGAGGCCAAUCAGUUUUCUUAAAUUGUAGAUACAU--UUAUAUGUGUUUGUGCUUUAAGAAUUAAAA ......((((((...(-(.....))....))).)))....((((((((.(..((((((((--....))))))))..).))))))))...... ( -20.90, z-score = -2.17, R) >droYak2.chr2R 19900114 89 - 21139217 GAAAGCGAUGGCAAAA-CUAUCGGUGGAGGCCAAUCAGUUUUCUUAAAUUGUAGAUACAU--UUAUAUGUGUUUGUGCUUUAAGAAUUAAAA ......((((((...(-(.....))....))).)))....((((((((.(..((((((((--....))))))))..).))))))))...... ( -20.90, z-score = -2.17, R) >droEre2.scaffold_4845 21296789 89 - 22589142 GAAAGCGAUGGCAAAA-CUAUCGGUGGAGGCCAAUCAGUUUUCUUAAAUUGUAGAUACAU--UUAUAUGUGUUUGUGCUUUAAGAAUUAAAA ......((((((...(-(.....))....))).)))....((((((((.(..((((((((--....))))))))..).))))))))...... ( -20.90, z-score = -2.17, R) >droAna3.scaffold_13266 5792244 83 - 19884421 ------GAAAGCAAAA-CUAUCGGUGGAGGCCAAUCAGUUUUCUUAAAUUGUAGAUACAU--UUAUAUGUGUUUGUGCUUUAAGAAUUGAAA ------..........-.....(((....)))..(((((..(((((((.(..((((((((--....))))))))..).)))))))))))).. ( -17.80, z-score = -1.64, R) >dp4.chr3 7607857 88 - 19779522 GAAAGCGAUGGC-AAA-CUAUCGGCAGGGGCCAAUCCGUUUUCUUGAAAUGUAGAUACAU--UUAUAUGUGUUUGUGGCUUAAGAAUUGAAA (((((((((((.-...-)))))(((....))).....))))))((((..(..((((((((--....))))))))..)..))))......... ( -21.90, z-score = -1.45, R) >droPer1.super_2 7803473 88 - 9036312 GAAAGCGAUGGC-AAA-CUAUCGGCAGGGGCCAAUCCGUUUUCUUGAAAUGUAGAUACAU--UUAUAUGUGUUUGUGGCUUAAGAAUUGAAA (((((((((((.-...-)))))(((....))).....))))))((((..(..((((((((--....))))))))..)..))))......... ( -21.90, z-score = -1.45, R) >droWil1.scaffold_180700 5037036 90 - 6630534 -GAGUCGAUAACAAAAGUUAUCAGUAGCGGUCCAUCUAUUUUC-UGAAUUGUAGAUACAUAAGUGUGUGUGUUUCUGCUUAAAGAAUUUAAA -.....((((((....))))))..............((..(((-(.....(((((.(((((......))))).)))))....))))..)).. ( -18.60, z-score = -1.73, R) >consensus GAAAGCGAUGGCAAAA_CUAUCGGUGGAGGCCAAUCAGUUUUCUUAAAUUGUAGAUACAU__UUAUAUGUGUUUGUGCUUUAAGAAUUAAAA ......(((((......)))))(((....)))........((((((((.(..((((((((......))))))))..).))))))))...... (-17.44 = -17.22 + -0.22)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:47:09 2011