
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 19,897,302 – 19,897,394 |
| Length | 92 |
| Max. P | 0.938553 |

| Location | 19,897,302 – 19,897,394 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 63.67 |
| Shannon entropy | 0.55276 |
| G+C content | 0.55667 |
| Mean single sequence MFE | -32.95 |
| Consensus MFE | -16.74 |
| Energy contribution | -17.68 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.855269 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
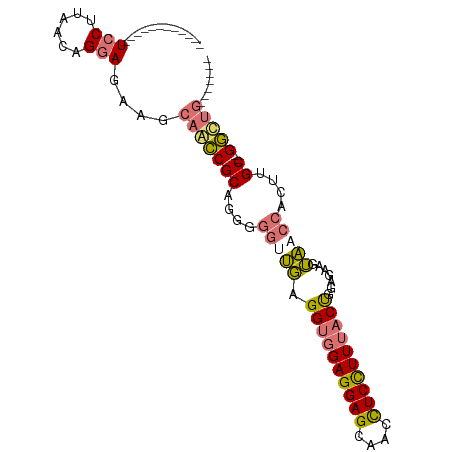
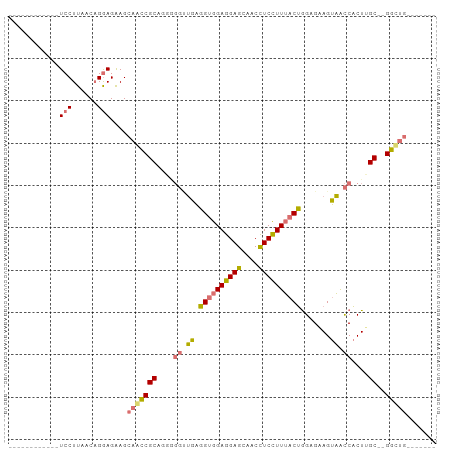
>dm3.chr2R 19897302 92 + 21146708 GAAGGAGUUACCUCCUUAGCAGGAGAAGCAACCGCACGGGGUUGAGGUGGAGGAGCAACCUCUUUUACUGGUGAAGUAACCACUUGC--GGUUG------- .((((((....))))))...........((((((((.(.(((((.((((((((((....)))))))))).......))))).).)))--)))))------- ( -34.20, z-score = -2.09, R) >droGri2.scaffold_15252 3023931 101 + 17193109 GGAGGAGUCGCUUCCUUAACCGGAGGAGGAGUCGCUUCCUUGACCGGAGGAGGAGUCGCUUCCUUGACCAGAGGAGGUGUCGCUUGCUUGACUGGAGGAGU (((((.(.(.(((((((.....))))))).).).)))))....((((((.(((.(.(((((((((.....))))))))).).))).))...))))...... ( -39.50, z-score = 0.00, R) >droYak2.chr2R 19875745 80 + 21139217 ------------UCCUUAACAGAAGAAGCGACCGCUGGAGGCUGAGGUGGAGGAGCAACCUCCUUUACUGGAGAAGUAACCACUUGC--GGCGG------- ------------................((.(((((((..(((..((((((((((....)))))))))).....)))..)))...))--)))).------- ( -27.00, z-score = -1.21, R) >droSim1.chr2R 18443173 80 + 19596830 ------------UCCUUAGCAGGAGAAGCAACCGCAGUGGGUUGAGGUGGAGGAGCAACCUCUUUUACUGGUGAAGUAACCACUGGC--GGUUG------- ------------((((....))))....((((((((((((.(((.((((((((((....))))))))))..))).....))))).))--)))))------- ( -31.10, z-score = -2.33, R) >consensus ____________UCCUUAACAGGAGAAGCAACCGCAGGGGGUUGAGGUGGAGGAGCAACCUCCUUUACUGGAGAAGUAACCACUUGC__GGCUG_______ ...........((((......))))...(((((((....(((((.((((((((((....)))))))))).......)))))....))..)))))....... (-16.74 = -17.68 + 0.94)


| Location | 19,897,302 – 19,897,394 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 63.67 |
| Shannon entropy | 0.55276 |
| G+C content | 0.55667 |
| Mean single sequence MFE | -28.00 |
| Consensus MFE | -12.29 |
| Energy contribution | -12.48 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.938553 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
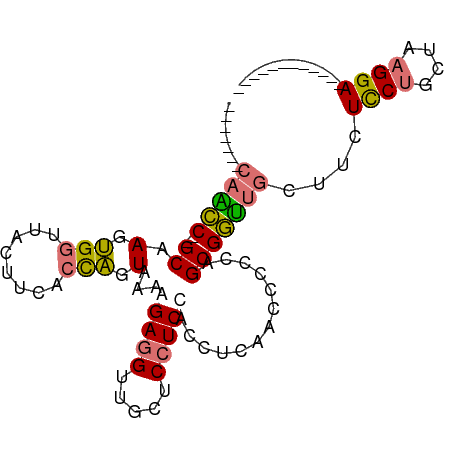
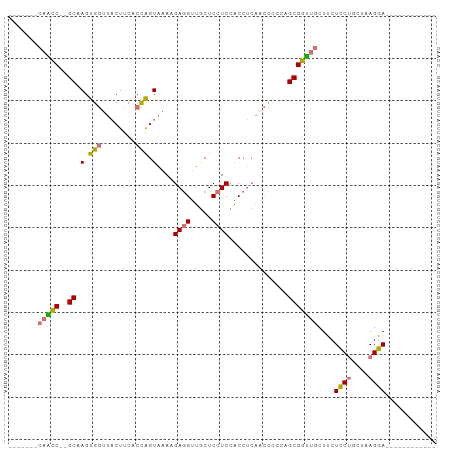
>dm3.chr2R 19897302 92 - 21146708 -------CAACC--GCAAGUGGUUACUUCACCAGUAAAAGAGGUUGCUCCUCCACCUCAACCCCGUGCGGUUGCUUCUCCUGCUAAGGAGGUAACUCCUUC -------(((((--(((.(.(((((((.....)))....(((((.(.....).))))))))).).))))))))..((((((....)))))).......... ( -29.90, z-score = -2.91, R) >droGri2.scaffold_15252 3023931 101 - 17193109 ACUCCUCCAGUCAAGCAAGCGACACCUCCUCUGGUCAAGGAAGCGACUCCUCCUCCGGUCAAGGAAGCGACUCCUCCUCCGGUUAAGGAAGCGACUCCUCC .........(((........)))(((......)))..((((..((..((((...((((...((((......))))...))))...))))..))..)))).. ( -29.00, z-score = -0.19, R) >droYak2.chr2R 19875745 80 - 21139217 -------CCGCC--GCAAGUGGUUACUUCUCCAGUAAAGGAGGUUGCUCCUCCACCUCAGCCUCCAGCGGUCGCUUCUUCUGUUAAGGA------------ -------.((((--((.(.(((........))).)...((((((((...........)))))))).)))).))....((((....))))------------ ( -23.70, z-score = -1.02, R) >droSim1.chr2R 18443173 80 - 19596830 -------CAACC--GCCAGUGGUUACUUCACCAGUAAAAGAGGUUGCUCCUCCACCUCAACCCACUGCGGUUGCUUCUCCUGCUAAGGA------------ -------(((((--((.((((((((((.....)))))..(((((.(.....).)))))...))))))))))))....((((....))))------------ ( -29.40, z-score = -3.89, R) >consensus _______CAACC__GCAAGUGGUUACUUCACCAGUAAAAGAGGUUGCUCCUCCACCUCAACCCCCAGCGGUUGCUUCUCCUGCUAAGGA____________ .......(((((..((.(.(((........))).)...(((((.....))))).............)))))))....((((....))))............ (-12.29 = -12.48 + 0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:47:07 2011