| Sequence ID | dm3.chr2R |
|---|---|
| Location | 19,682,146 – 19,682,247 |
| Length | 101 |
| Max. P | 0.873382 |

| Location | 19,682,146 – 19,682,247 |
|---|---|
| Length | 101 |
| Sequences | 9 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 73.73 |
| Shannon entropy | 0.51532 |
| G+C content | 0.45304 |
| Mean single sequence MFE | -28.20 |
| Consensus MFE | -11.94 |
| Energy contribution | -11.86 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.732804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 19682146 101 + 21146708 CCUGUUGGCUGGAAAAGGGCGAGCGAAUCUUUUGGCCAUUGUUUU-------AUAAUGAUAAAACAAUGAAUGCGG--UCUGGGAACCUGCACCUUGAGUCGCUAGUCCG-- ((........))....((((.(((((..(((..((.(((((((((-------((....)))))))))))..(((((--(......))).)))))..)))))))).)))).-- ( -33.20, z-score = -2.50, R) >droSim1.chr2R 18249276 97 + 19596830 ----CCUGCUGGAAAAGGGCGAGCGAAUCUCUCGGCCAUUGUUUU-------AUAAUGAUAAAACAAUGAAUGCGG--UCUGGGAACCUGCACCUUGAGUCGCUAGUGCG-- ----(((........)))((.(((((..(((..((.(((((((((-------((....)))))))))))..(((((--(......))).)))))..))))))))...)).-- ( -30.50, z-score = -1.92, R) >droSec1.super_9 2972292 97 + 3197100 ----CCUGCUGGAAAAGGGCGAGCGAAUCUCUCGGCCAUUGUUUU-------AUAAUGAUAAAACAAUGAAUGCGG--UCUGGGAACCUGCACCUUGAGUCGCUAGUCCG-- ----((....))....((((.(((((..(((..((.(((((((((-------((....)))))))))))..(((((--(......))).)))))..)))))))).)))).-- ( -34.10, z-score = -3.18, R) >droYak2.chr2R 19664105 98 + 21139217 ----CCUGUUGGAAAAGGGCGAGCGAAUCUCUCGACCAUUGUUUU-------AUAAUGAUAAAACAAUGAAUGCGG--UCUGGGAACCUGCACCUUGAGUCGCUCGUCGCU- ----((....)).....(((((((((....(((((.(((((((((-------((....)))))))))))..(((((--(......))).)))..))))))))))))))...- ( -34.80, z-score = -3.56, R) >droEre2.scaffold_4845 21042464 97 + 22589142 ----CCUGCUGGAAAAGGGCGAGCGAAUCUCUCGACCAUUGUUUU-------AUAAUGAUAAAACAAUGAAUGUGG--UCUGGGAACCUGCACCUUGAGUCGCUCAUAGU-- ----(((........)))..((((((....(((((.(((((((((-------((....))))))))))).....((--(.(((....))).)))))))))))))).....-- ( -29.80, z-score = -2.60, R) >droAna3.scaffold_13266 359773 96 + 19884421 ----GCUGA-GGAAAAGGGCGAGCGAAUCUUUCAGCCAUUGUUUU-------AUCAUUAUAAUACAAUGAAUGUGG--UUUGGGAACUUGCACCUUGAGCCAAGGGGCUG-- ----.....-....(((((((((((((((...((..((((((.((-------((....)))).))))))..)).))--))))....))))).)))).((((....)))).-- ( -27.40, z-score = -1.24, R) >dp4.chr3 19471562 93 - 19779522 -------UUUGGAUAAGGGCGAGCGAAUCUUUAGACCAUUGUCUUGAU---CAUUUUUAUUAGACAAUGGAUGUGUGCUUUGGGAACUUGUACCUUGAGUCGG--------- -------...(((((((..((((((..((....))((((((((((((.---.....)))..))))))))).....)))))...)..))))).)).........--------- ( -21.60, z-score = -0.57, R) >droPer1.super_2 8544033 93 - 9036312 -------UUUGGAUAAGGGCGAGCGAAUCUUUAGACCAUUGUCUUGAU---CAUUUUUAUUAGACAAUGGAUGUGUGCUUUGGGAACUUGUACCUUGAGUCGG--------- -------...(((((((..((((((..((....))((((((((((((.---.....)))..))))))))).....)))))...)..))))).)).........--------- ( -21.60, z-score = -0.57, R) >droWil1.scaffold_180745 974619 99 + 2843958 -------AGUAUAUAAGGGCGAGCGAAUCGUU--GCCAUUGUUUCCAAAAAUACAAACACGAAACAAUGAAUUCCG---UUGGGAACUUGUACCU-GCACCUUGAAUGUGUG -------..(((.(((((((((.....))))(--(((((((((((...............)))))))))..((((.---...)))).........-)))))))).))).... ( -20.76, z-score = -0.21, R) >consensus ____CCUGCUGGAAAAGGGCGAGCGAAUCUUUCGGCCAUUGUUUU_______AUAAUGAUAAAACAAUGAAUGCGG__UCUGGGAACCUGCACCUUGAGUCGCUAGUCCG__ ..........((..((((((((.....)).......(((((((((...............)))))))))....................)).))))...))........... (-11.94 = -11.86 + -0.08)



| Location | 19,682,146 – 19,682,247 |
|---|---|
| Length | 101 |
| Sequences | 9 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 73.73 |
| Shannon entropy | 0.51532 |
| G+C content | 0.45304 |
| Mean single sequence MFE | -23.87 |
| Consensus MFE | -9.69 |
| Energy contribution | -9.37 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.873382 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 19682146 101 - 21146708 --CGGACUAGCGACUCAAGGUGCAGGUUCCCAGA--CCGCAUUCAUUGUUUUAUCAUUAU-------AAAACAAUGGCCAAAAGAUUCGCUCGCCCUUUUCCAGCCAACAGG --.((.(.(((((.((..(((((.((((....))--)))))))(((((((((((....))-------))))))))).......)).))))).).))................ ( -28.20, z-score = -2.95, R) >droSim1.chr2R 18249276 97 - 19596830 --CGCACUAGCGACUCAAGGUGCAGGUUCCCAGA--CCGCAUUCAUUGUUUUAUCAUUAU-------AAAACAAUGGCCGAGAGAUUCGCUCGCCCUUUUCCAGCAGG---- --.((...((((((((..(((((.((((....))--))))...(((((((((((....))-------))))))))))))..)))..))))).))(((........)))---- ( -28.70, z-score = -2.90, R) >droSec1.super_9 2972292 97 - 3197100 --CGGACUAGCGACUCAAGGUGCAGGUUCCCAGA--CCGCAUUCAUUGUUUUAUCAUUAU-------AAAACAAUGGCCGAGAGAUUCGCUCGCCCUUUUCCAGCAGG---- --.((.(.((((((((..(((((.((((....))--))))...(((((((((((....))-------))))))))))))..)))..))))).).))............---- ( -30.70, z-score = -3.17, R) >droYak2.chr2R 19664105 98 - 21139217 -AGCGACGAGCGACUCAAGGUGCAGGUUCCCAGA--CCGCAUUCAUUGUUUUAUCAUUAU-------AAAACAAUGGUCGAGAGAUUCGCUCGCCCUUUUCCAACAGG---- -.....((((((((((...((((.((((....))--))))))((((((((((((....))-------))))))))))....)))..))))))).(((........)))---- ( -31.30, z-score = -3.57, R) >droEre2.scaffold_4845 21042464 97 - 22589142 --ACUAUGAGCGACUCAAGGUGCAGGUUCCCAGA--CCACAUUCAUUGUUUUAUCAUUAU-------AAAACAAUGGUCGAGAGAUUCGCUCGCCCUUUUCCAGCAGG---- --.....(((((((((........((((....))--))....((((((((((((....))-------))))))))))....)))..))))))..(((........)))---- ( -26.10, z-score = -2.41, R) >droAna3.scaffold_13266 359773 96 - 19884421 --CAGCCCCUUGGCUCAAGGUGCAAGUUCCCAAA--CCACAUUCAUUGUAUUAUAAUGAU-------AAAACAAUGGCUGAAAGAUUCGCUCGCCCUUUUCC-UCAGC---- --.((((....)))).((((.((.(((.......--...((..((((((.((((....))-------)).))))))..))........))).))))))....-.....---- ( -18.67, z-score = -0.50, R) >dp4.chr3 19471562 93 + 19779522 ---------CCGACUCAAGGUACAAGUUCCCAAAGCACACAUCCAUUGUCUAAUAAAAAUG---AUCAAGACAAUGGUCUAAAGAUUCGCUCGCCCUUAUCCAAA------- ---------.(((.((..((........))............(((((((((..........---....)))))))))......)).)))................------- ( -14.24, z-score = -1.19, R) >droPer1.super_2 8544033 93 + 9036312 ---------CCGACUCAAGGUACAAGUUCCCAAAGCACACAUCCAUUGUCUAAUAAAAAUG---AUCAAGACAAUGGUCUAAAGAUUCGCUCGCCCUUAUCCAAA------- ---------.(((.((..((........))............(((((((((..........---....)))))))))......)).)))................------- ( -14.24, z-score = -1.19, R) >droWil1.scaffold_180745 974619 99 - 2843958 CACACAUUCAAGGUGC-AGGUACAAGUUCCCAA---CGGAAUUCAUUGUUUCGUGUUUGUAUUUUUGGAAACAAUGGC--AACGAUUCGCUCGCCCUUAUAUACU------- .........((((.((-..((....(((.((((---((((((.....)))))))..........(((....)))))).--))).....))..)))))).......------- ( -22.70, z-score = -1.13, R) >consensus __CGGACUAGCGACUCAAGGUGCAGGUUCCCAGA__CCGCAUUCAUUGUUUUAUCAUUAU_______AAAACAAUGGCCGAAAGAUUCGCUCGCCCUUUUCCAGCAGG____ ..........(((.((..((........))............((((((((((...............))))))))))......)).)))....................... ( -9.69 = -9.37 + -0.32)


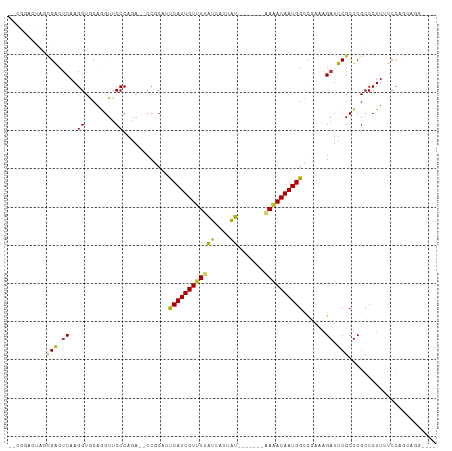
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:46:36 2011