
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 19,654,425 – 19,654,524 |
| Length | 99 |
| Max. P | 0.637511 |

| Location | 19,654,425 – 19,654,524 |
|---|---|
| Length | 99 |
| Sequences | 8 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 85.48 |
| Shannon entropy | 0.27324 |
| G+C content | 0.37028 |
| Mean single sequence MFE | -24.25 |
| Consensus MFE | -17.09 |
| Energy contribution | -16.61 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.637511 |
| Prediction | RNA |
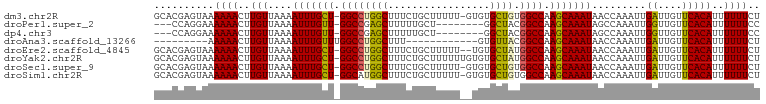
Download alignment: ClustalW | MAF
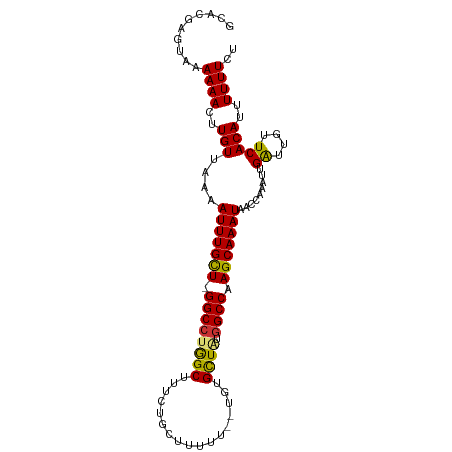
>dm3.chr2R 19654425 99 + 21146708 GCACGAGUAAAAAACUUGUUAAAAUUUGCU-GGCCUGGCUUUCUGCUUUUU-GUGUGCUGUGGCCAAGCAAAUAACCAAAUUGAUUGUUCACAUUUUUUCU ..((((((.....))))))....(((((((-((((((((...(........-)...)))).)))).)))))))........(((....))).......... ( -22.80, z-score = -0.90, R) >droPer1.super_2 8511211 89 - 9036312 ---CCAGGAAAAAACUUGUUAAAAUUUGUU-GGCCGAGCUUUUUGCU--------GGCUACGGCCAAGCAAAUAGCCAAAUUGGUUGUUCACAUUUUUUCC ---...((((((((..((.....(((((((-(((((((((.......--------)))).))))).)))))))((((.....))))...))..)))))))) ( -29.80, z-score = -3.20, R) >dp4.chr3 19439995 89 - 19779522 ---CCAGGAAAAAACUUGUUAAAAUUUGUU-GGCCGAGCUUUUUGCU--------GGCUACGGCCAAGCAAAUAGCCAAAUUGGUUGUUCACAUUUUUUCC ---...((((((((..((.....(((((((-(((((((((.......--------)))).))))).)))))))((((.....))))...))..)))))))) ( -29.80, z-score = -3.20, R) >droAna3.scaffold_13266 333340 80 + 19884421 ---------AAAAACUUGUUAAAAUUUGUUUGGCCUGGCUUU------------GUGUUACGGCCAAGCAAAUAACCAAAUUGAUUGUUCACAUUUUUUCU ---------.((((..(((....((((((((((((((((...------------..)))).)))))))))))).........((....)))))..)))).. ( -20.70, z-score = -2.73, R) >droEre2.scaffold_4845 21014965 98 + 22589142 GCACGAGUAAAAAACUUGUUAAAAUUUGCU-GGCCUGGCUUUCUGCUUUUU--UGUGCUAUGGCCAAGCAAAUAACCAAAUUGAUUGUUCACAUUUUUUCU ..((((((.....))))))....(((((((-((((((((............--...)))).)))).)))))))........(((....))).......... ( -23.36, z-score = -1.62, R) >droYak2.chr2R 19632423 100 + 21139217 GCACGAGUAAAAAACUUGUUAAAAUUUGCU-GGCCUGGCUUUCUGCUUUUUUGUGUGCUAUGGCCAAGCAAAUAACCAAAUUGAUUGUUCACAUUUUUUCU ..((((((.....))))))....(((((((-((((((((....(((......))).)))).)))).)))))))........(((....))).......... ( -23.80, z-score = -1.54, R) >droSec1.super_9 2944805 99 + 3197100 GCACGAGUAAAAAACUUGUUAAAAUUUGCU-GGCCUGGCUUUCUGCUUUUU-GUGUGCUGUGGCCAAGCAAAUAACCAAAUUGAUUGUUCACAUUUUUUCU ..((((((.....))))))....(((((((-((((((((...(........-)...)))).)))).)))))))........(((....))).......... ( -22.80, z-score = -0.90, R) >droSim1.chr2R 18221648 99 + 19596830 GCACGAGUAAAAAACUUGUUAAAAUUUGCU-GGCAUGGCUUUCUGCUUUUU-GUGUGCUGUGGCCAAGCAAAUAACCAAAUUGAUUGUUCACAUUUUUUCU ..((((((.....))))))....(((((((-((((((((...(........-)...))))).))).)))))))........(((....))).......... ( -20.90, z-score = -0.07, R) >consensus GCACGAGUAAAAAACUUGUUAAAAUUUGCU_GGCCUGGCUUUCUGCUUUUU__UGUGCUAUGGCCAAGCAAAUAACCAAAUUGAUUGUUCACAUUUUUUCU ..........((((..(((....(((((((.((((((((.................)))).)))).))))))).........((....)))))..)))).. (-17.09 = -16.61 + -0.48)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:46:34 2011