
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 19,597,316 – 19,597,437 |
| Length | 121 |
| Max. P | 0.856636 |

| Location | 19,597,316 – 19,597,437 |
|---|---|
| Length | 121 |
| Sequences | 7 |
| Columns | 128 |
| Reading direction | forward |
| Mean pairwise identity | 63.45 |
| Shannon entropy | 0.67800 |
| G+C content | 0.47985 |
| Mean single sequence MFE | -35.56 |
| Consensus MFE | -12.34 |
| Energy contribution | -11.01 |
| Covariance contribution | -1.34 |
| Combinations/Pair | 1.67 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.856636 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 19597316 121 + 21146708 UUGCGCCUCCCAUCGAUAGCAGUCC-AACUGGAUUU-UGCAUGCUAUCGAUAUUUUCCCAAUUUAAAUUUGUAGUUCUAACAGC-GAGCACCAGUGGCGUCUUCCGCUUCCGGUUGUUGCUUGG---- .(((.......((((((((((((((-....))))..-....)))))))))).................((((.((....)).))-)))))((((..(((....(((....))).)))..).)))---- ( -31.40, z-score = -1.43, R) >droSim1.chr2R 18163367 121 + 19596830 UUGCGCCUCCCACCGAUAGCAGUCC-AAUUGGAUUU-UGCAUGCUAUCGAUACUUUCCCAAUUUAAAUUUUUAGUUCUAACAGC-GAGCACCAGUGGCGCCUUCCGUUUCCGGUGCUUGCUUGG---- .........(((.((((((((((((-....))))..-....))))))))................................(((-(((((((.(..(((.....)))..).)))))))))))))---- ( -36.10, z-score = -3.08, R) >droSec1.super_9 2876318 121 + 3197100 UUGCGCCUCCCACCGAUAGCAGUCC-AAUUGGAUUU-UGCAUGCUAUCGAUACUUUCCCAAUUUUAAUUUUUAGUUCUAACAGC-GAGCACCAGUGGCGCCUUCCGUUUCCGGUGCUUGCUUGG---- .........(((.((((((((((((-....))))..-....))))))))................................(((-(((((((.(..(((.....)))..).)))))))))))))---- ( -36.10, z-score = -3.17, R) >droYak2.chr2R 19574164 121 + 21139217 UUGCGCCUCCCACCGAUAGCAGUCCCAACUGGAUUU-UGCAUGCUAUCGAUAUUUUUAUAAUUCAAAUUUAUAGUUAUAAAAGC-AACCAUGCGGGGCGCCA-CCGCUUCCGGUUAUUGCUUUA---- ..(((((((....((((((((((((.....))))..-....))))))))....((((((((((.........))))))))))((-(....)))))))))).(-(((....))))..........---- ( -34.50, z-score = -2.61, R) >droEre2.scaffold_4845 20959158 120 + 22589142 UUGCGCCUCCCACCGAUAGCGGUGC-AACUGGUUUU-UGCAUGCUAUCGAUAUUUUUGCAAUUUGAAUUUGUAGUACUAAAAGC-GGGCACGCGUGGCGCCA-CCGUUUCCGGUUGUUGCUUGA---- ..(((...(((..((((((((.(((-((.......)-))))))))))))......((((((.......))))))..........-)))..)))(..(((..(-(((....)))))))..)....---- ( -36.50, z-score = -0.32, R) >dp4.chr3 10375364 127 - 19779522 UCGU-UCAUACAACGAACGUGUUGCGAGCGAACUUUCUGCGUACGCUCCCAGGUUCUUCGGCUCUAGCGUUCGGUUCGACUCGUUGUGUGCCGGUUGUGGGCACCAUACUUUACACGUGCAUAUUUUA ((((-(.....)))))((((((.((((((((((......((.(((((....((..(....)..))))))).))))))).)))))((.(((((.......)))))))......)))))).......... ( -38.20, z-score = -0.50, R) >droPer1.super_4 2436315 123 - 7162766 UCGU-UCAUACAACGAACGUGUUGCGAGCGAACUUUCUGCGUACGCUCCCAGGUUCUUCGGCUCUAGCGUUCGGCUCGACUCGUUGUGUGCCGGUUGUGUAC----UUUUUUACACGUGCAUAUUUUA (((.-.((((((((((.((.((..(((((((((((...((....))....))))))....((....))))))))).))..)))))))))).))).(((((((----..........)))))))..... ( -36.10, z-score = -0.72, R) >consensus UUGCGCCUCCCACCGAUAGCAGUCC_AACUGGAUUU_UGCAUGCUAUCGAUAUUUUUCCAAUUUAAAUUUUUAGUUCUAACAGC_GAGCACCAGUGGCGCCU_CCGUUUCCGGUUCUUGCUUGG____ ..(((((.(....((((((((....................))))))))........................((.((........)).))..).)))))............................ (-12.34 = -11.01 + -1.34)


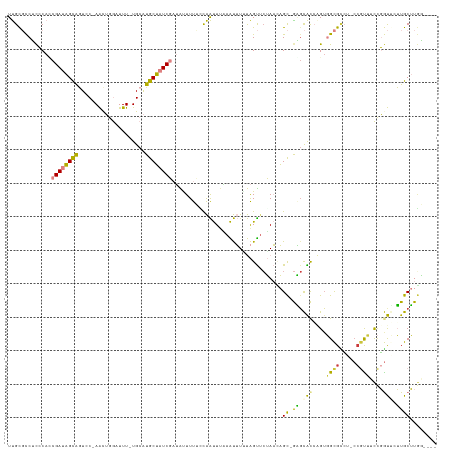
| Location | 19,597,316 – 19,597,437 |
|---|---|
| Length | 121 |
| Sequences | 7 |
| Columns | 128 |
| Reading direction | reverse |
| Mean pairwise identity | 63.45 |
| Shannon entropy | 0.67800 |
| G+C content | 0.47985 |
| Mean single sequence MFE | -36.05 |
| Consensus MFE | -14.35 |
| Energy contribution | -14.70 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.841550 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 19597316 121 - 21146708 ----CCAAGCAACAACCGGAAGCGGAAGACGCCACUGGUGCUC-GCUGUUAGAACUACAAAUUUAAAUUGGGAAAAUAUCGAUAGCAUGCA-AAAUCCAGUU-GGACUGCUAUCGAUGGGAGGCGCAA ----....((....(((((..(((.....)))..)))))((((-.(..(((((........)))))...).))...((((((((((((.((-(.......))-).).)))))))))))....)))).. ( -32.90, z-score = -1.18, R) >droSim1.chr2R 18163367 121 - 19596830 ----CCAAGCAAGCACCGGAAACGGAAGGCGCCACUGGUGCUC-GCUGUUAGAACUAAAAAUUUAAAUUGGGAAAGUAUCGAUAGCAUGCA-AAAUCCAAUU-GGACUGCUAUCGGUGGGAGGCGCAA ----...........(((....)))...((((((((....(((-(...(((((........)))))..))))..)))(((((((((((.((-(.......))-).).))))))))))....))))).. ( -36.90, z-score = -1.62, R) >droSec1.super_9 2876318 121 - 3197100 ----CCAAGCAAGCACCGGAAACGGAAGGCGCCACUGGUGCUC-GCUGUUAGAACUAAAAAUUAAAAUUGGGAAAGUAUCGAUAGCAUGCA-AAAUCCAAUU-GGACUGCUAUCGGUGGGAGGCGCAA ----...........(((....)))...(((((.(..((....-....(((....)))........))..).....((((((((((((.((-(.......))-).).)))))))))))...))))).. ( -35.79, z-score = -1.42, R) >droYak2.chr2R 19574164 121 - 21139217 ----UAAAGCAAUAACCGGAAGCGG-UGGCGCCCCGCAUGGUU-GCUUUUAUAACUAUAAAUUUGAAUUAUAAAAAUAUCGAUAGCAUGCA-AAAUCCAGUUGGGACUGCUAUCGGUGGGAGGCGCAA ----..........((((....)))-).(((((((..((((((-(......)))))))..................(((((((((((.(..-...(((.....))))))))))))))))).))))).. ( -37.90, z-score = -2.09, R) >droEre2.scaffold_4845 20959158 120 - 22589142 ----UCAAGCAACAACCGGAAACGG-UGGCGCCACGCGUGCCC-GCUUUUAGUACUACAAAUUCAAAUUGCAAAAAUAUCGAUAGCAUGCA-AAAACCAGUU-GCACCGCUAUCGGUGGGAGGCGCAA ----....((....((((....)))-).)).....((((.(((-((.....(((..............)))........(((((((.((((-(.......))-)))..))))))))))))..)))).. ( -40.24, z-score = -2.39, R) >dp4.chr3 10375364 127 + 19779522 UAAAAUAUGCACGUGUAAAGUAUGGUGCCCACAACCGGCACACAACGAGUCGAACCGAACGCUAGAGCCGAAGAACCUGGGAGCGUACGCAGAAAGUUCGCUCGCAACACGUUCGUUGUAUGA-ACGA ..........((((((......(((((((.......))))).)).((((.(((((((.(((((....(((.......))).))))).))......)))))))))..))))))(((((.....)-)))) ( -36.50, z-score = -0.70, R) >droPer1.super_4 2436315 123 + 7162766 UAAAAUAUGCACGUGUAAAAAA----GUACACAACCGGCACACAACGAGUCGAGCCGAACGCUAGAGCCGAAGAACCUGGGAGCGUACGCAGAAAGUUCGCUCGCAACACGUUCGUUGUAUGA-ACGA .......(((..(((((.....----.)))))..((((....(..((..((.(((.....))).))..))..)...))))(((((.((.......)).))))))))...(((((((...))))-))). ( -32.10, z-score = -0.21, R) >consensus ____CCAAGCAAGAACCGGAAACGG_AGGCGCCACUGGUGCUC_GCUGUUAGAACUAAAAAUUAAAAUUGGAAAAAUAUCGAUAGCAUGCA_AAAUCCAGUU_GGACUGCUAUCGGUGGGAGGCGCAA ........((.....(((....))).....(((..((((..............))))...................((((((((((......................))))))))))...))))).. (-14.35 = -14.70 + 0.36)


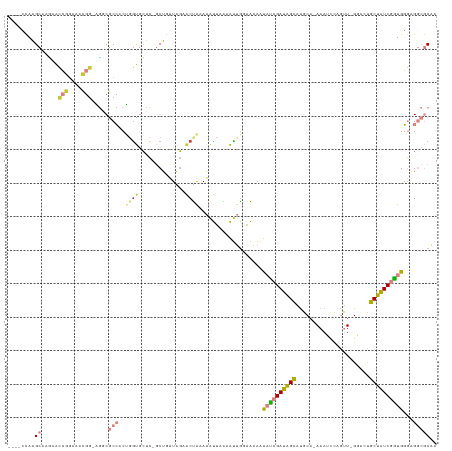
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:46:16 2011