
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 19,269,014 – 19,269,078 |
| Length | 64 |
| Max. P | 0.648759 |

| Location | 19,269,014 – 19,269,078 |
|---|---|
| Length | 64 |
| Sequences | 12 |
| Columns | 66 |
| Reading direction | forward |
| Mean pairwise identity | 73.24 |
| Shannon entropy | 0.58825 |
| G+C content | 0.49778 |
| Mean single sequence MFE | -10.48 |
| Consensus MFE | -7.38 |
| Energy contribution | -7.18 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.40 |
| Mean z-score | -0.29 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.648759 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 19269014 64 + 21146708 --UUUGCUCUCUCAUUUAGAUGGCGAUGCCACACCCUCGAUAUUCGUGACCAUUCCGCGUUUUGCC --.(((((.(((.....))).)))))...........(((....((((.......))))..))).. ( -8.50, z-score = 0.61, R) >droSim1.chr2R 17852372 64 + 19596830 --UUUGCUCUCUAAUUCAGAUGGCGAUGCCACGCCCUCGAUAUUCGUGACCAUUCCGCGUUUCGCC --...((...........((.((((......)))).))((....((((.......))))..)))). ( -13.10, z-score = -0.43, R) >droSec1.super_9 2563470 64 + 3197100 --UUUGCUCUCUAAUUCAGAUGGCGAUGCCACGCCCUCGAUAUUCGUGACCAUUCCGCGUUUCGCC --...((...........((.((((......)))).))((....((((.......))))..)))). ( -13.10, z-score = -0.43, R) >droYak2.chr2R 19237865 62 + 21139217 --UUUGCUCUUC--CUUAGAUGGCGAUGCCACGCCCUCGAUAUUUGUGACCAUUCCGCGUUUCGCC --...((.....--....((.((((......)))).))((....((((.......))))..)))). ( -11.40, z-score = -0.41, R) >droEre2.scaffold_4845 20639221 62 + 22589142 --UUUGCUCUCC--UUUAGAUGGCGAUGCCACGCCCUCGAUAUUUGUGACCAUUCCGCGUUUCGCC --...((.....--....((.((((......)))).))((....((((.......))))..)))). ( -11.40, z-score = -0.41, R) >droAna3.scaffold_13266 11988434 62 - 19884421 ----CCCAUUUCUUUCCAGAUGGUGAUCAGACACCUUCAAUUUUCGUGACUAUUCCCCGGUUUGCC ----............(((((((.((..((.(((...........))).))..)).)).))))).. ( -7.10, z-score = 0.89, R) >dp4.chr3 9135494 62 + 19779522 ----AUCCCUUUUGUUCAGAUGGCGAUCCAACGCCCUCAAUCUUUGUGACCAUUCCGCGCUUUGCC ----..............((.((((......)))).)).......(((.......)))........ ( -9.20, z-score = -0.21, R) >droPer1.super_4 4457477 62 + 7162766 ----AUCCCUUUUGUUCAGAUGGCGAUCCAACGCCCUCAAUCUUUGUGACCAUUCCGCGCUUUGCC ----..............((.((((......)))).)).......(((.......)))........ ( -9.20, z-score = -0.21, R) >droWil1.scaffold_181141 4283797 62 + 5303230 ----UAUCUUAUACUGCAGAUGGGGAUCCUACACCAUCUAUUUUUGUGACCAUUCCACGUUUUGCC ----...........(((((((((.......).))))))......(((.......))).....)). ( -10.40, z-score = -0.58, R) >droVir3.scaffold_12875 18411460 62 + 20611582 --GUUUCCUUCUU-U-UAGAUGGUGACAAGACGCCCUCCAUCUUUGUGACCAUACCGCGUUUCACU --...........-.-...(((((.((((((.(.....).)).)))).)))))............. ( -9.60, z-score = -0.66, R) >droMoj3.scaffold_6496 12061810 62 + 26866924 --CUUUCUCUCGC-U-UAGAUGGUGACAAAACACCCUCGAUCUUUGUGACUAUACCUCGCUUCACU --.......((((-.-.((((((((......))))....))))..))))................. ( -10.90, z-score = -2.07, R) >droGri2.scaffold_15245 4524667 65 + 18325388 CUCUCUCUCUCGG-UACAGAUGGUGACAAAACGCCGUCGAUCUUUAUGACCAUUCCACGAUUCGGC .........(((.-....(((((((......)))))))(.((.....)).)......)))...... ( -11.90, z-score = 0.43, R) >consensus __UUUGCUCUCUU_UUCAGAUGGCGAUGCCACGCCCUCGAUAUUUGUGACCAUUCCGCGUUUCGCC ..................((.((((......)))).))......((((.......))))....... ( -7.38 = -7.18 + -0.19)

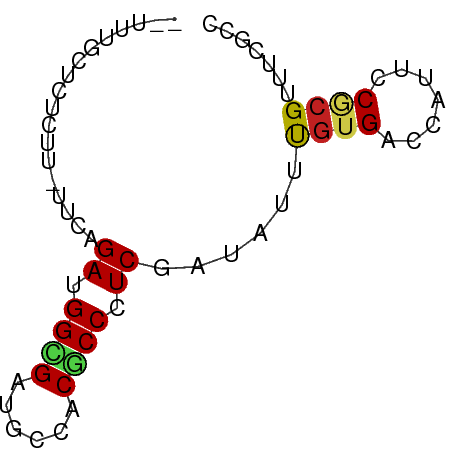

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:45:27 2011