
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 3,736,787 – 3,736,872 |
| Length | 85 |
| Max. P | 0.983321 |

| Location | 3,736,787 – 3,736,872 |
|---|---|
| Length | 85 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 61.34 |
| Shannon entropy | 0.67192 |
| G+C content | 0.37411 |
| Mean single sequence MFE | -13.69 |
| Consensus MFE | -8.48 |
| Energy contribution | -8.84 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.08 |
| Mean z-score | -0.88 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.74 |
| SVM RNA-class probability | 0.964527 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
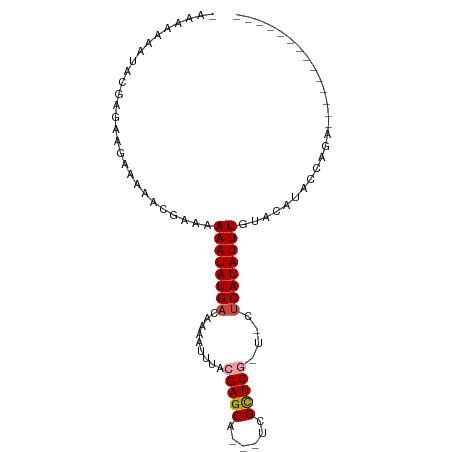
>dm3.chr2L 3736787 85 + 23011544 -AAUAACAGAUGAGAAGAAAAGCGAAAAAAUAUGACAAAAUUUACGAGCA---UCGCUCGUUCCUCAUAUUUGUACAUACCAGACUCUC--------- -..........((((............((((((((........((((((.---..))))))...)))))))).............))))--------- ( -13.41, z-score = -1.30, R) >droPer1.super_1 3798758 97 - 10282868 -AAGGGUAUACGUGAAAAAACAACGUAAAAUAUGACAAAAUUUACGAGCAGCAUCGCUCGCUUCUCAUAUUUGUACAUACCAGAGGACUCCACUUCUC -...(((((((((.........)))))((((((((.........(((((......)))))....)))))))).....))))(((((......))))). ( -21.52, z-score = -2.52, R) >dp4.chr4_group3 6699944 97 - 11692001 -AAGGGUAUACGUGAAAAAACAACGUAAAAUAUGACAAAAUUUACGAGCAGCAUCGCUCGCUUCUCAUAUUUGUACAUACCAGAGGACUCCACUUCUC -...(((((((((.........)))))((((((((.........(((((......)))))....)))))))).....))))(((((......))))). ( -21.52, z-score = -2.52, R) >droAna3.scaffold_12916 5802301 73 + 16180835 -----------GUGAAGAAAAGCGAAAAAAUAUGACAAAAUUUACGAGCA---UCGCUCGUUCCUCAUAUUUGUACAGACCAGACUC----------- -----------................((((((((........((((((.---..))))))...))))))))...............----------- ( -11.30, z-score = -0.68, R) >droVir3.scaffold_12963 5230904 69 - 20206255 GGAAAAAGUUGAACGUGAAAGACGAGAAAAUAUGACAAAAUUUACGAGCG---CUGUUCA---CACAUAUUUGGA----------------------- ..............(((((((.((.(.((((........)))).)...))---)).))))---)...........----------------------- ( -7.00, z-score = 0.83, R) >droGri2.scaffold_15126 4406118 69 + 8399593 GGAAAAAGUUGAACGUGAAAGACGAGAAAAUAUGACAAAAUUUACGAGCG---CAGCUCA---CUCAUAUUUGUA----------------------- .............(((.....)))...((((((((..........((((.---..)))).---.))))))))...----------------------- ( -7.40, z-score = 0.90, R) >consensus _AAAAAAAUACGAGAAGAAAAACGAAAAAAUAUGACAAAAUUUACGAGCA___UCGCUCG_U_CUCAUAUUUGUACAUACCAGA______________ ...........................((((((((.........(((((......)))))....)))))))).......................... ( -8.48 = -8.84 + 0.36)


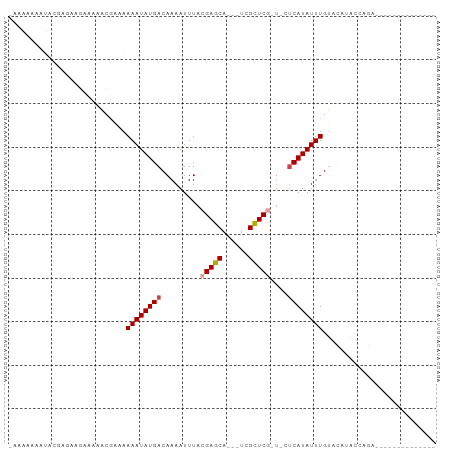
| Location | 3,736,787 – 3,736,872 |
|---|---|
| Length | 85 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 61.34 |
| Shannon entropy | 0.67192 |
| G+C content | 0.37411 |
| Mean single sequence MFE | -16.27 |
| Consensus MFE | -10.27 |
| Energy contribution | -11.17 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.13 |
| SVM RNA-class probability | 0.983321 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
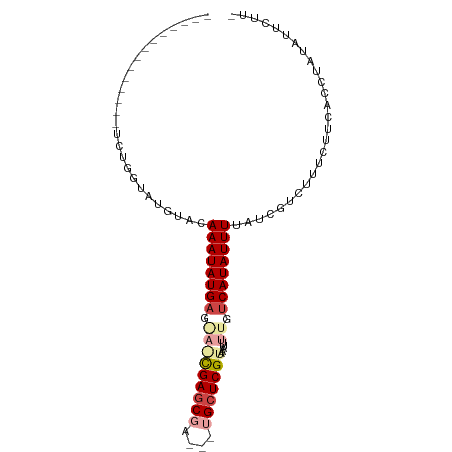
>dm3.chr2L 3736787 85 - 23011544 ---------GAGAGUCUGGUAUGUACAAAUAUGAGGAACGAGCGA---UGCUCGUAAAUUUUGUCAUAUUUUUUCGCUUUUCUUCUCAUCUGUUAUU- ---------(((((...(((......((((((((.((((((((..---.)))))).....)).))))))))....)))....)))))..........- ( -18.80, z-score = -1.33, R) >droPer1.super_1 3798758 97 + 10282868 GAGAAGUGGAGUCCUCUGGUAUGUACAAAUAUGAGAAGCGAGCGAUGCUGCUCGUAAAUUUUGUCAUAUUUUACGUUGUUUUUUCACGUAUACCCUU- .....((((((....(....(((((.((((((((.(((((((((....))))))).....)).))))))))))))).)...))))))..........- ( -23.60, z-score = -1.38, R) >dp4.chr4_group3 6699944 97 + 11692001 GAGAAGUGGAGUCCUCUGGUAUGUACAAAUAUGAGAAGCGAGCGAUGCUGCUCGUAAAUUUUGUCAUAUUUUACGUUGUUUUUUCACGUAUACCCUU- .....((((((....(....(((((.((((((((.(((((((((....))))))).....)).))))))))))))).)...))))))..........- ( -23.60, z-score = -1.38, R) >droAna3.scaffold_12916 5802301 73 - 16180835 -----------GAGUCUGGUCUGUACAAAUAUGAGGAACGAGCGA---UGCUCGUAAAUUUUGUCAUAUUUUUUCGCUUUUCUUCAC----------- -----------(((...(((......((((((((.((((((((..---.)))))).....)).))))))))....)))...)))...----------- ( -14.70, z-score = -0.71, R) >droVir3.scaffold_12963 5230904 69 + 20206255 -----------------------UCCAAAUAUGUG---UGAACAG---CGCUCGUAAAUUUUGUCAUAUUUUCUCGUCUUUCACGUUCAACUUUUUCC -----------------------...........(---((((..(---((.....((((........))))...)))..))))).............. ( -7.00, z-score = -0.62, R) >droGri2.scaffold_15126 4406118 69 - 8399593 -----------------------UACAAAUAUGAG---UGAGCUG---CGCUCGUAAAUUUUGUCAUAUUUUCUCGUCUUUCACGUUCAACUUUUUCC -----------------------.(((((((((((---((.....---))))))))...))))).................................. ( -9.90, z-score = -1.01, R) >consensus ______________UCUGGUAUGUACAAAUAUGAG_A_CGAGCGA___UGCUCGUAAAUUUUGUCAUAUUUUAUCGUCUUUCUUCACCUAUAUUCUU_ ..........................((((((((.((((((((......)))))).....)).))))))))........................... (-10.27 = -11.17 + 0.89)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:14:31 2011