
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 19,225,032 – 19,225,124 |
| Length | 92 |
| Max. P | 0.653161 |

| Location | 19,225,032 – 19,225,124 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 78.57 |
| Shannon entropy | 0.40162 |
| G+C content | 0.45582 |
| Mean single sequence MFE | -20.84 |
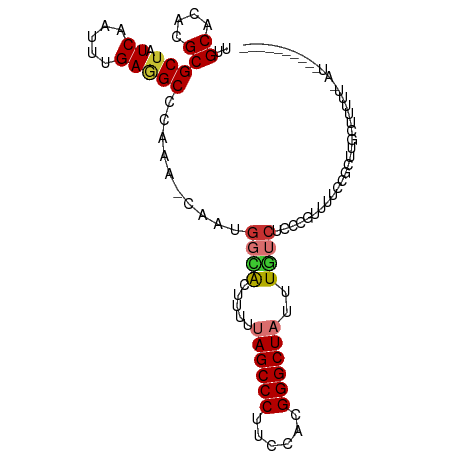
| Consensus MFE | -12.32 |
| Energy contribution | -12.60 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.653161 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
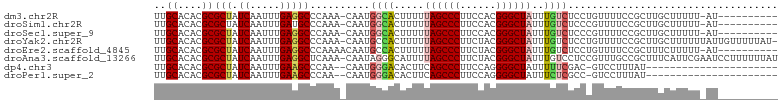
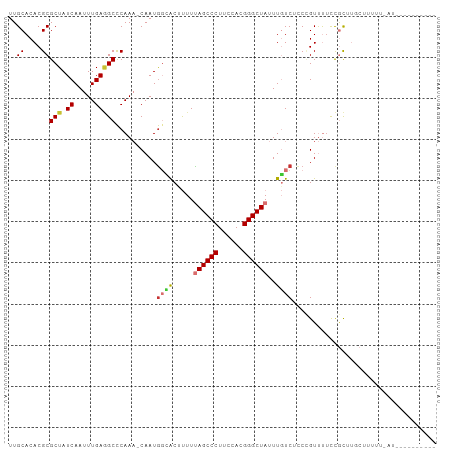
>dm3.chr2R 19225032 92 + 21146708 UUGCACACGCGCUAUCAAUUUGAGGCCCAAA-CAAUGGCACUUUUUAGCCCUUCCACGGGCUAUUUGUCUCCUGUUUUCCGCUUGCUUUUU-AU---------- ..(((...(((.....(((..((((((((..-...))).......((((((......))))))...)))))..)))...))).))).....-..---------- ( -21.40, z-score = -1.42, R) >droSim1.chr2R 17808126 92 + 19596830 UUGCACACGCGCUAUCAAUUUGAUGCCCAAA-CAAUGGCACUUUUUAGCCCUUCCACGGGCUAUUUGUCUCCCGUUUUCCGCUUGCUUUUU-AU---------- ..(((...((((.(((.....))))).....-.(((((.((....((((((......))))))...))...)))))....)).))).....-..---------- ( -19.70, z-score = -1.48, R) >droSec1.super_9 2517960 92 + 3197100 UUGCACACGCGCUAUCAAUUUGAGGCCCAAA-CAAUGGCACUUUUUAGCCCUUCCACGGGCUAUUUGUCUCCCGUUUUCCGCUUGCUUUUU-AU---------- ..(((...(((.....(((..((((((((..-...))).......((((((......))))))...)))))..)))...))).))).....-..---------- ( -20.50, z-score = -1.08, R) >droYak2.chr2R 19191010 102 + 21139217 UUGCACACGCGCUAUCAAUUUGAGGCCCAAA-CAAUGCCACUUUUUAGCCCUUCUACGGGCUAUUUGUCUCCUGUUUUCCGCUUGCUUUUUUAUUGUUUUUAU- ..(((...(((.....(((..(((((.((..-...))........((((((......))))))...)))))..)))...))).))).................- ( -18.10, z-score = -0.63, R) >droEre2.scaffold_4845 20595561 93 + 22589142 UUGCACACGCGCUAUCAAUUUGAGGCCCAAAACAAUGCCACUUUUUAGCCCUUCUACGGGCUAUUUGUCUCCUGUUUUCCGCUUUCUUUUU-AU---------- ..((....))...........(((((..((((((.....((....((((((......))))))...))....))))))..)))))......-..---------- ( -18.30, z-score = -1.54, R) >droAna3.scaffold_13266 8320221 103 - 19884421 UUGCACACGCGCUAUCAAUUUGAGGCUCAAA-CAAUAGGGCAUUUUAGCCCUUCUACGGGCUAUUUGUCCUCCGUUUGCCGCUUUCAUUCGAAUCCUUUUUUAU ..((....)).......(((((((((.((((-(...((((((...((((((......))))))..))))))..)))))..))).....)))))).......... ( -28.00, z-score = -2.92, R) >dp4.chr3 16736921 79 - 19779522 UUGCACACGCGCUAUCAAUUUGAAGCCCAA--CAAUGGGACACUUCAGCCCUUCCAGGGGCUAUUUUUCGAC-GUCCUUUAU---------------------- ......(((((..........((((((((.--...))))...))))((((((....))))))......)).)-)).......---------------------- ( -19.40, z-score = -1.64, R) >droPer1.super_2 5800042 79 - 9036312 UUGCACACGCGCUAUCAAUUUGAAGCCCAA--CAAUGGGACACUUCAGCCCUUCCAGGGGCUAUUUCUCGCC-GUCCUUUAU---------------------- ......(((.((.........((((((((.--...))))...))))((((((....)))))).......)))-)).......---------------------- ( -21.30, z-score = -1.88, R) >consensus UUGCACACGCGCUAUCAAUUUGAGGCCCAAA_CAAUGGCACUUUUUAGCCCUUCCACGGGCUAUUUGUCUCCCGUUUUCCGCUUGCUUUUU_AU__________ ..((....))(((.((.....)))))..........((((.....((((((......))))))..))))................................... (-12.32 = -12.60 + 0.28)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:45:21 2011