| Sequence ID | dm3.chr2R |
|---|---|
| Location | 19,146,694 – 19,146,789 |
| Length | 95 |
| Max. P | 0.756600 |

| Location | 19,146,694 – 19,146,789 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 81.37 |
| Shannon entropy | 0.35404 |
| G+C content | 0.56152 |
| Mean single sequence MFE | -34.14 |
| Consensus MFE | -18.70 |
| Energy contribution | -19.90 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.517077 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 19146694 95 + 21146708 CAAGAAAAGCGGGCCAAGAGGGCCAAGCCAUAUAAGAGACCGC--GAUGAUGGCGGCAAUUAUACGUGGUGACCAUGUGGCGGGUGGGGU--UUUCCAG ...(((((.(.((((.....))))..(((((((.....(((((--((((((.......))))).))))))....)))))))......).)--))))... ( -32.00, z-score = -1.58, R) >droSim1.chrU 6598511 95 + 15797150 CAAGAAAAGCGGGCCAAGAGGGCCAAGCCAUAUAAGAGCCCGC--GAUGAUGGCGGCAAUUAUACGUGGUGGCCAUGUGGCGGGUGGGGU--UUUCCAG ...(((((.(.((((.....)))).............((((((--.((.(((((.((...........)).))))))).))))))..).)--))))... ( -34.90, z-score = -1.37, R) >droSec1.super_9 2440130 95 + 3197100 CAAGAAAAGCGGGCCAAGAGGGCCAAGCCAUAUAAGAGCCCGC--GAUGAUGGCGGCAAUUAUACGUGGUGGCCAUGUGGCGGGUGGGGU--UUUCCAG ...(((((.(.((((.....)))).............((((((--.((.(((((.((...........)).))))))).))))))..).)--))))... ( -34.90, z-score = -1.37, R) >droYak2.chr2R 19111864 96 + 21139217 CAAGAAAAGCGGGCCAAGAGGGCCAAGCCAUAUAAGAGUCUGC--GAUGAUGGCGGCAAUUAUACGUGGUGGCCAUGUGGCGGGCGGGGU-UUUUCCAC ...(((((((.((((.....)))).............((((((--.((.(((((.((...........)).))))))).))))))...))-)))))... ( -34.40, z-score = -1.72, R) >droEre2.scaffold_4845 20514958 97 + 22589142 CAAGAAAAGCGGGCCAAGAGGGCCAAGCCAUAUAAGAGCCCGC--GAUGAUGGCGGCAAUUAUACGUGGUGGCCAUGUGGCGGGUGGUGUGUUUUCCAC ...(((((((.((((.....))))..))(((((....((((((--.((.(((((.((...........)).))))))).)))))).))))))))))... ( -37.50, z-score = -2.03, R) >droMoj3.scaffold_6496 4839742 84 - 26866924 CAACAAUGGCCGGC-----AGACCAAAAUGUGUAAACACGCGUAUGAUGAUGGCGGCAAUUAUACGGUCAGCACGAGAUGCUGGCCGGA---------- ........((((.(-----(.......((((((....)))))).......)).)))).......(((((((((.....)))))))))..---------- ( -31.14, z-score = -2.85, R) >consensus CAAGAAAAGCGGGCCAAGAGGGCCAAGCCAUAUAAGAGCCCGC__GAUGAUGGCGGCAAUUAUACGUGGUGGCCAUGUGGCGGGUGGGGU__UUUCCAG ........((..((((....((((..(((((((((..(.((((.........)))))..)))))..))))))))...))))..)).............. (-18.70 = -19.90 + 1.20)
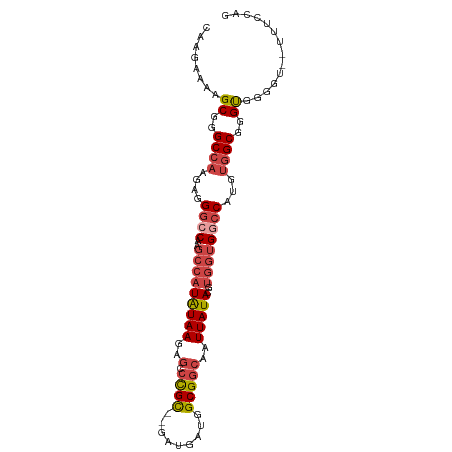



| Location | 19,146,694 – 19,146,789 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 81.37 |
| Shannon entropy | 0.35404 |
| G+C content | 0.56152 |
| Mean single sequence MFE | -27.32 |
| Consensus MFE | -18.19 |
| Energy contribution | -19.47 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.756600 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 19146694 95 - 21146708 CUGGAAA--ACCCCACCCGCCACAUGGUCACCACGUAUAAUUGCCGCCAUCAUC--GCGGUCUCUUAUAUGGCUUGGCCCUCUUGGCCCGCUUUUCUUG .(((...--...)))...((((...(((((...(((((((..(((((.......--)))))...)))))))...)))))....))))............ ( -26.20, z-score = -1.81, R) >droSim1.chrU 6598511 95 - 15797150 CUGGAAA--ACCCCACCCGCCACAUGGCCACCACGUAUAAUUGCCGCCAUCAUC--GCGGGCUCUUAUAUGGCUUGGCCCUCUUGGCCCGCUUUUCUUG .(((...--...)))...((((...(((((...(((((((..(((((.......--)).)))..)))))))...)))))....))))............ ( -28.80, z-score = -1.66, R) >droSec1.super_9 2440130 95 - 3197100 CUGGAAA--ACCCCACCCGCCACAUGGCCACCACGUAUAAUUGCCGCCAUCAUC--GCGGGCUCUUAUAUGGCUUGGCCCUCUUGGCCCGCUUUUCUUG .(((...--...)))...((((...(((((...(((((((..(((((.......--)).)))..)))))))...)))))....))))............ ( -28.80, z-score = -1.66, R) >droYak2.chr2R 19111864 96 - 21139217 GUGGAAAA-ACCCCGCCCGCCACAUGGCCACCACGUAUAAUUGCCGCCAUCAUC--GCAGACUCUUAUAUGGCUUGGCCCUCUUGGCCCGCUUUUCUUG ((((....-...))))..((((...(((((...(((((((..((.((.......--)).).)..)))))))...)))))....))))............ ( -24.70, z-score = -1.01, R) >droEre2.scaffold_4845 20514958 97 - 22589142 GUGGAAAACACACCACCCGCCACAUGGCCACCACGUAUAAUUGCCGCCAUCAUC--GCGGGCUCUUAUAUGGCUUGGCCCUCUUGGCCCGCUUUUCUUG ((((........))))..((((...(((((...(((((((..(((((.......--)).)))..)))))))...)))))....))))............ ( -31.10, z-score = -2.26, R) >droMoj3.scaffold_6496 4839742 84 + 26866924 ----------UCCGGCCAGCAUCUCGUGCUGACCGUAUAAUUGCCGCCAUCAUCAUACGCGUGUUUACACAUUUUGGUCU-----GCCGGCCAUUGUUG ----------..(((.((((((...)))))).))).(((((.((((.((..((((.....(((....)))....)))).)-----).)))).))))).. ( -24.30, z-score = -1.47, R) >consensus CUGGAAA__ACCCCACCCGCCACAUGGCCACCACGUAUAAUUGCCGCCAUCAUC__GCGGGCUCUUAUAUGGCUUGGCCCUCUUGGCCCGCUUUUCUUG ..................((((...(((((...(((((((..(((((.........)))).)..)))))))...)))))....))))............ (-18.19 = -19.47 + 1.28)
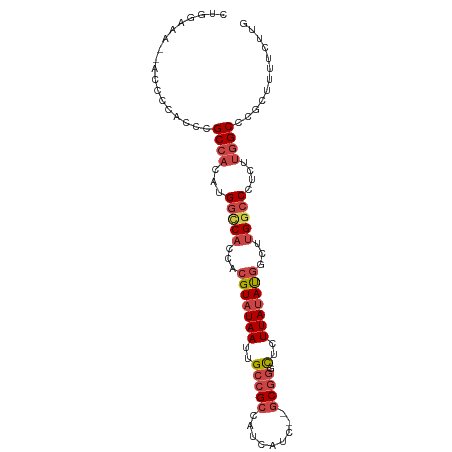



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:45:15 2011