
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 3,686,501 – 3,686,600 |
| Length | 99 |
| Max. P | 0.648645 |

| Location | 3,686,501 – 3,686,600 |
|---|---|
| Length | 99 |
| Sequences | 8 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 73.68 |
| Shannon entropy | 0.49392 |
| G+C content | 0.50457 |
| Mean single sequence MFE | -24.08 |
| Consensus MFE | -14.79 |
| Energy contribution | -15.04 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.02 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.648645 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 3686501 99 + 23011544 UCUAUCGGUGGGCGGUCUCAUCAG-ACGGCAAAGAGGCCAUGGGUGGGCAGAACAAAAACAAAAAAAAAAACCGGGAAUAAAAAGUAUU-----GGCUCACAUGU .......((((((.(((.((((..-..(((......)))...)))))))...........................((((.....))))-----.)))))).... ( -20.00, z-score = -0.42, R) >droSim1.chr2L 3643929 91 + 22036055 UCUGUCGGUGGGCGGUCUCAUCAG-ACGGCAAAGAGGCCAUGGGUGGGCAGAUU-----CC---GAAAAAAACGGGAAUAAAAAGUAUU-----GGCUCACAUGU .......((((((.(((.((((..-..(((......)))...)))))))...((-----((---(.......)))))............-----.)))))).... ( -25.70, z-score = -1.27, R) >droSec1.super_5 1782097 94 + 5866729 UCUAUCGGUGGGCGGUCUCAUCAG-ACGGCAAAGAGGCCAUGGGUGGGCAGAUU-----CCCAAAAAAAAAACGGGAAUAAAAAGUAUU-----GGCUCACAUGU .......((((((.(((.((((..-..(((......)))...)))))))..(((-----(((...........))))))..........-----.)))))).... ( -27.90, z-score = -2.23, R) >droYak2.chr2L 3680404 96 + 22324452 UCUAUCGGUGGGCGGUCUCAUCAGGACGGCAAAGAGGCCAUGGGUGGGCAGCAAA-----AAAGAAUAAAAGUGGGUAUAAAAAGUAUUU----GGCUCACAUGU ((((((.((((((.((((.....)))).)).......)))).)))))).......-----...........((((((...(((....)))----.)))))).... ( -23.71, z-score = -1.62, R) >droEre2.scaffold_4929 3723206 85 + 26641161 UCCAUCGGUGGGCGGUCUCAUCAG-ACGGCAAAGAGGCCAUGGGUGGGCAGA--------------AAAAAGUGGGAAUAAAAAGUAUU-----GGCUCACAUGU ((((((.((((((.((((....))-)).)).......)))).))))))....--------------.....(((((.(((.....))).-----..))))).... ( -23.61, z-score = -1.48, R) >droAna3.scaffold_12916 5742903 96 + 16180835 UUUAUCGGUGGGCGGUCUCAUCAG-AUGGCAAAGAGGCCACGGGAGGGCCGACA------CAAAAAAAAAAAUAAAACUGGAAUAGAUUCUU--GGCUCACAUGU .......(((((((((((..((.(-.((((......)))))))..)))))....------.................(.((((....)))).--))))))).... ( -24.50, z-score = -1.57, R) >dp4.chr4_group3 6648599 91 - 11692001 UGUAUCAGUGGGCGGUCUCAUCAG-CCAGCAAAGAGGCCACGGGCGGCCCAAC-------------AAAGACUCGACCCCGAAGUCCUCAACUCCACUCACAUGU (((.....(((((.((((.....(-((........)))...)))).)))))..-------------...((((((....)).)))).............)))... ( -23.60, z-score = 0.22, R) >droPer1.super_1 3748131 91 - 10282868 UGUAUCAGUGGGCGGUCUCAUCAG-CCAGCAAAGAGGCCACGGGCGGCCCAAC-------------AAAGACUCGACCCCGAAGUCCUCAACUCCACUCACAUGU (((.....(((((.((((.....(-((........)))...)))).)))))..-------------...((((((....)).)))).............)))... ( -23.60, z-score = 0.22, R) >consensus UCUAUCGGUGGGCGGUCUCAUCAG_ACGGCAAAGAGGCCAUGGGUGGGCAGAC___________AAAAAAAAUGGGAAUAAAAAGUAUU_____GGCUCACAUGU .......((((((((((((..............))))))..(......)..............................................)))))).... (-14.79 = -15.04 + 0.25)

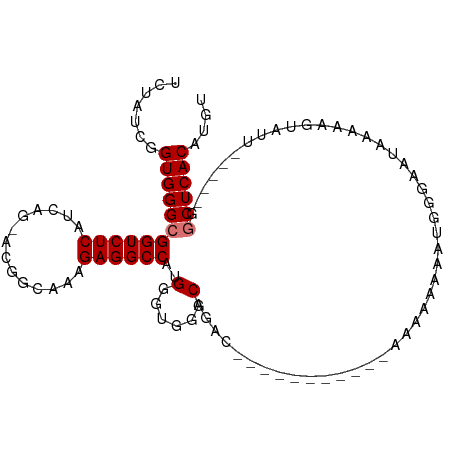
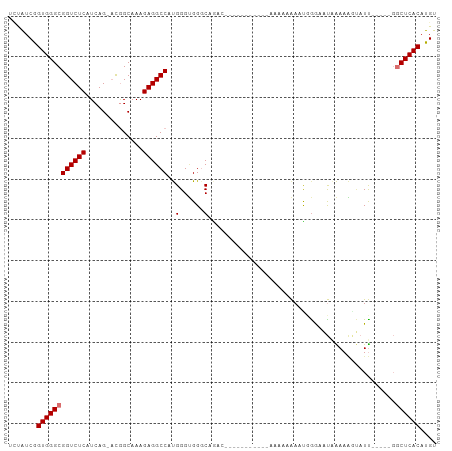
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:14:27 2011