| Sequence ID | dm3.chr2R |
|---|---|
| Location | 18,898,174 – 18,898,268 |
| Length | 94 |
| Max. P | 0.933783 |

| Location | 18,898,174 – 18,898,268 |
|---|---|
| Length | 94 |
| Sequences | 10 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 73.23 |
| Shannon entropy | 0.49668 |
| G+C content | 0.39591 |
| Mean single sequence MFE | -24.86 |
| Consensus MFE | -13.06 |
| Energy contribution | -13.61 |
| Covariance contribution | 0.55 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.38 |
| SVM RNA-class probability | 0.933783 |
| Prediction | RNA |
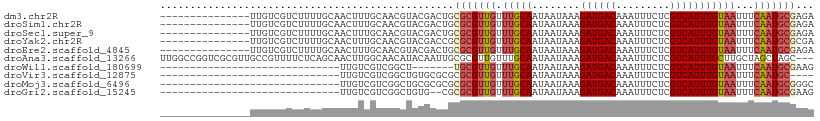
Download alignment: ClustalW | MAF
>dm3.chr2R 18898174 94 - 21146708 ---------------UUGUCGUCUUUUGCAACUUUGCAACGUACGACUGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGCGAGA ---------------((((((((((((((((....((((((..(....)..))))))))))).....)))))))))))...((((((...(((......))).)))))) ( -30.60, z-score = -3.56, R) >droSim1.chr2R 17486346 94 - 19596830 ---------------UUGUCGUCUUUUGCAACUUUGCAACGUACGACUGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGCGAGA ---------------((((((((((((((((....((((((..(....)..))))))))))).....)))))))))))...((((((...(((......))).)))))) ( -30.60, z-score = -3.56, R) >droSec1.super_9 2190083 94 - 3197100 ---------------UUGUCGUCUUUUGCAACUUUGCAACGUACGACUGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGCGAGA ---------------((((((((((((((((....((((((..(....)..))))))))))).....)))))))))))...((((((...(((......))).)))))) ( -30.60, z-score = -3.56, R) >droYak2.chr2R 18859017 94 - 21139217 ---------------UUGUCGUCUUUUGCAACUUUGCAACGUACGACCGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGCGCGA ---------------..(((((.(.(((((....))))).).)))))(((((((((.(((((........((((((.........)))))))))))...))))))))). ( -31.60, z-score = -3.71, R) >droEre2.scaffold_4845 20259904 94 - 22589142 ---------------UUGUCGUCUUUUGCAACUUUGCAACGUACGACUGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGCGAGA ---------------((((((((((((((((....((((((..(....)..))))))))))).....)))))))))))...((((((...(((......))).)))))) ( -30.60, z-score = -3.56, R) >droAna3.scaffold_13266 15234312 106 - 19884421 UUGGCCGGUCGCGUUGCCGUUUUCUCAGCAACUUGGCAACAUACAAUUGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGCUUGCUAGCUAGC--- (((((..((.(((((((((..............))))))).....((((((.......)))))).....(((((((.........)))))))))..))..))))).--- ( -26.74, z-score = -0.09, R) >droWil1.scaffold_180699 1977038 72 + 2593675 ------------------------------UUGUCGUCGGCU-------UGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGCGAAG ------------------------------((((((((...(-------((((.....))))).......)))))))).....((((...(((......))).)))).. ( -15.70, z-score = -1.20, R) >droVir3.scaffold_12875 7068983 75 - 20611582 ------------------------------UUGUCGUCGGCUGUGCGCGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGC---- ------------------------------((((((((....(((....)))((((....))))......))))))))...........................---- ( -14.30, z-score = 1.01, R) >droMoj3.scaffold_6496 25778645 79 - 26866924 ------------------------------UUGUCGUCGGCUGCGCGCGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGCGGGC ------------------------------((((((((....(((....)))((((....))))......))))))))....(((((...(((......))).))))). ( -20.90, z-score = -0.08, R) >droGri2.scaffold_15245 616575 77 - 18325388 ------------------------------UUGUCGUCGGCUGUG--CGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGCGAAG ------------------------------((((((((.((....--.))(((.....))).........)))))))).....((((...(((......))).)))).. ( -17.00, z-score = -0.08, R) >consensus _______________UUGUCGUCUUUUGCAACUUUGCAACGUACGACUGCGCGUUGUUUGCAAUAAUAAAGAUGACAAAUUUCUCGUCAUUUGUAAUUUCAAUGCGAGA .................................................(((((((.(((((........((((((.........)))))))))))...)))))))... (-13.06 = -13.61 + 0.55)
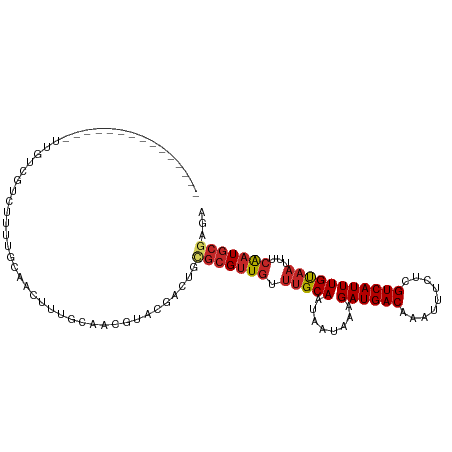
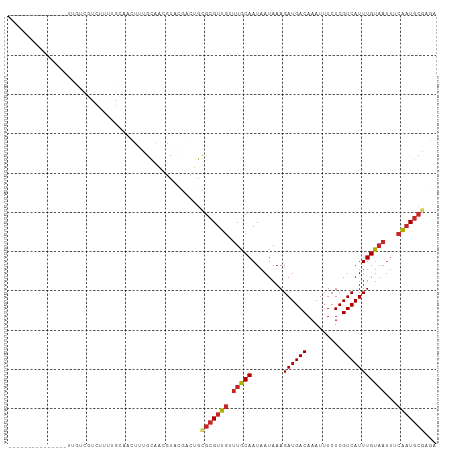
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:44:37 2011