
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 18,403,184 – 18,403,286 |
| Length | 102 |
| Max. P | 0.647405 |

| Location | 18,403,184 – 18,403,286 |
|---|---|
| Length | 102 |
| Sequences | 7 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 72.98 |
| Shannon entropy | 0.54202 |
| G+C content | 0.51249 |
| Mean single sequence MFE | -30.39 |
| Consensus MFE | -16.89 |
| Energy contribution | -18.22 |
| Covariance contribution | 1.33 |
| Combinations/Pair | 1.25 |
| Mean z-score | -0.96 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.647405 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 18403184 102 + 21146708 AGUGCCGGAACAAAGGCAGCCUUCAUGAUAUCCUCGGAUGAGCAUUUGGCUGGUAU-GCAGCUUCAUUCUCAUGGGUUGCUUCUGGU----UCCCGUCCUCAUUCCU ((..(.(((((..((((((((..(((((.......((((((......(((((....-.))))))))))))))))))))))))...))----))).)..))....... ( -33.01, z-score = -1.10, R) >droSim1.chr2R 16995980 106 + 19596830 AGUGCCGGAACAAAGGCAGCCUUCAUGAUAUCCUCGGAUGAGCAUUUGGCUGGUAU-GCAGCUUCAUUCUCAUGCGCUGCUUCUGGUCUUCUUCCUUCCUCAUUCCU ...((((((.....((((((......((.....))(.(((((.((..(((((....-.)))))..)).))))).))))))))))))).................... ( -29.20, z-score = -0.34, R) >droSec1.super_9 1711499 102 + 3197100 AGUGCCGGAACAAAGGCAGCCUUCAUGAUAUCCUCGGAUGAGCAUUUGGCUGCUAU-GCAGCUUCAUUCUCAUGCGCUGCUUCUGGU----UCCCCUCCUCAUACCU ......(((((..(((((((......((.....))(.(((((.((..((((((...-))))))..)).))))).))))))))...))----)))............. ( -33.20, z-score = -1.70, R) >droYak2.chr2R 15873403 98 + 21139217 AGUGGCGGAACAAAGACAGCCUUCAUGAUAUCCUCGGAUGAGCAUUUGGCUGGUAU-GCAGCUUCAUUCUCAUGCGCUGCUUCUGGU----UCCGCUGCCCCU---- .(..(((((((..((((((((...(((.((((....))))..)))..)))))....-(((((..(((....))).))))).))).))----)))))..)....---- ( -38.30, z-score = -2.52, R) >droEre2.scaffold_4845 15046228 98 - 22589142 AGUGCCGGAACAAAGACAGCCUUCAUGAUAUCCUCGGAUGAGCAUUUGGCUGGUAU-GCAGCUUCAUUCUCAUGGGCUGCUUCUGGU----UCCCCUCUUCCU---- .(.(((((((......(((((...(((.((((....))))..)))..)))))....-((((((.(((....))))))))))))))))----.)..........---- ( -30.70, z-score = -0.84, R) >droWil1.scaffold_180745 235915 90 + 2843958 --------UGUGUGGCAAGCCUUCAUGAUAUCCUCGGAUGAGCAUUUGGCUGGUAU-GCGCCGCCCGCCGUUCUCGAUGCCAUUCCUCUUUGCCUAUAU-------- --------((((.((((((.......((.....))(((((.(((((((((.(((..-.....))).)))).....))))))))))...)))))))))).-------- ( -23.80, z-score = 0.17, R) >droVir3.scaffold_12875 12515029 93 + 20611582 AGCGCCUCAACAAAACGAACUUUCAGAAUAUUCUCGGAUGAGCAUUUGGCUGGUGUUGCAGCUUCUGCUUCUU---CUUCUUC-GGUUCGGUGCCGU---------- .(((((..........((....)).((((......(((.(((((...(((((......)))))..)))))..)---)).....-.)))))))))...---------- ( -24.50, z-score = -0.41, R) >consensus AGUGCCGGAACAAAGGCAGCCUUCAUGAUAUCCUCGGAUGAGCAUUUGGCUGGUAU_GCAGCUUCAUUCUCAUGCGCUGCUUCUGGU____UCCCCUCCUCAU____ ...(((((((......(((((...(((.((((....))))..)))..))))).....(((((.............)))))))))))).................... (-16.89 = -18.22 + 1.33)


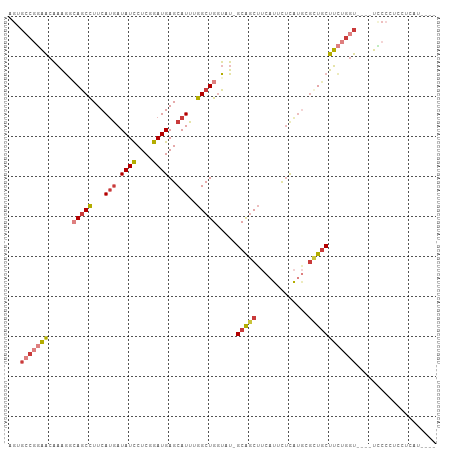
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:43:31 2011