
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 18,158,015 – 18,158,114 |
| Length | 99 |
| Max. P | 0.851724 |

| Location | 18,158,015 – 18,158,114 |
|---|---|
| Length | 99 |
| Sequences | 8 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 85.35 |
| Shannon entropy | 0.25893 |
| G+C content | 0.37508 |
| Mean single sequence MFE | -21.39 |
| Consensus MFE | -14.54 |
| Energy contribution | -14.91 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.851724 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 18158015 99 + 21146708 ------------------AUACACACGUACAACUGGGGCAUUAUCCCAAAACGAAAUGACAUCCCUUGGAAAUUAAUCCUUAAUUCCAUUCAUUUCGUAUAAUCCCAGAGAUAUACU ------------------..............(((((..(((((......(((((((((.......(((((.(((.....)))))))).)))))))))))))))))))......... ( -19.40, z-score = -2.21, R) >droSim1.chr2R 16757697 99 + 19596830 ------------------ACACACACGUACAGCUGGGGCAUUAUCCCAAAACGAAAUGACAUCCCUUGGAAAUUAAUCCUUAAUUCCAUUCAUUUCGUAUAAUCCCAGAGAUAUACU ------------------........(((.(.(((((..(((((......(((((((((.......(((((.(((.....)))))))).)))))))))))))))))))...).))). ( -19.90, z-score = -2.30, R) >droSec1.super_9 1468236 99 + 3197100 ------------------ACACACACGUACAGCUGGGGCAUUAUCCCAAAACGAAAUGACAUCCCUUGGAAAUUAAUCCUUAAUUCCAUUCAUUUCGUAUAAUCCCAGAGAUAUACU ------------------........(((.(.(((((..(((((......(((((((((.......(((((.(((.....)))))))).)))))))))))))))))))...).))). ( -19.90, z-score = -2.30, R) >droYak2.chr2R 17908060 99 + 21139217 ------------------ACACACACGUACAGCUGGGGCAUUAUCCCAAAACGAAAUGACAUCCCUUGGAAAUUAAUCCUUAAUUCCAUUCAUUUCGUAUAAUCCCAGAGAUAUACU ------------------........(((.(.(((((..(((((......(((((((((.......(((((.(((.....)))))))).)))))))))))))))))))...).))). ( -19.90, z-score = -2.30, R) >droEre2.scaffold_4845 12299048 98 + 22589142 ------------------ACACACACGUACAGCUGGG-CAUUAUCCCAAAACGAAAUGACAUCCCUUGGAAAUUAAUCCUUAAUUCCAUUCAUUUCGUAUAAUCCCAGAGAUAUACU ------------------........(((.(.(((((-.(((((......(((((((((.......(((((.(((.....)))))))).)))))))))))))))))))...).))). ( -21.80, z-score = -3.69, R) >droAna3.scaffold_13266 2167087 100 + 19884421 -----------------GACACACACGUAGUGGCAGGGCAUUAUCCCAAAACGAAAUGACAUCCCUUGGAAAUUAAUCCUUAAUUCCAUUCAUUUCGUAUAAUCCCAGAGAUAUACU -----------------....(((.....)))...(((.(((((......(((((((((.......(((((.(((.....)))))))).)))))))))))))))))........... ( -19.40, z-score = -1.60, R) >dp4.chr3 4799826 117 - 19779522 ACACAAAUGUAUUUGUAUCUGUGUGCAGGGAGAGAGAGCAUUAUCCCAAAACGAAAUGACAUCCCUUGGAAAUUAAUCCUUAAUUCCAUUCAUUUCGUAUAAUCCCAGAGAUAUACU .............(((((((.((....((((............))))...(((((((((.......(((((.(((.....)))))))).))))))))).......)).))))))).. ( -25.40, z-score = -1.73, R) >droPer1.super_4 4072080 117 + 7162766 ACACAAAUGUAUUUGUAUCUGUGUGCAGGGAGAGAGAGCAUUAUCCCAAAACGAAAUGACAUCCCUUGGAAAUUAAUCCUUAAUUCCAUUCAUUUCGUAUAAUCCCAGAGAUAUACU .............(((((((.((....((((............))))...(((((((((.......(((((.(((.....)))))))).))))))))).......)).))))))).. ( -25.40, z-score = -1.73, R) >consensus __________________ACACACACGUACAGCUGGGGCAUUAUCCCAAAACGAAAUGACAUCCCUUGGAAAUUAAUCCUUAAUUCCAUUCAUUUCGUAUAAUCCCAGAGAUAUACU ...................................(((.(((((......(((((((((.......(((((.(((.....)))))))).)))))))))))))))))........... (-14.54 = -14.91 + 0.37)
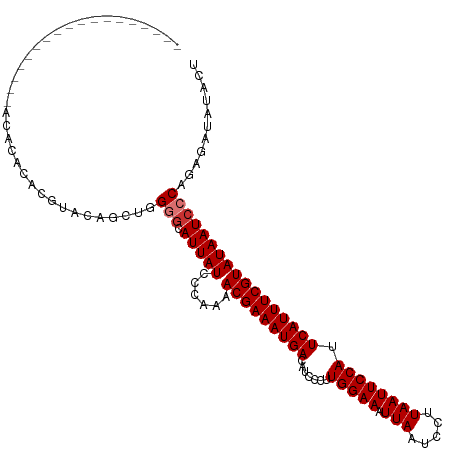


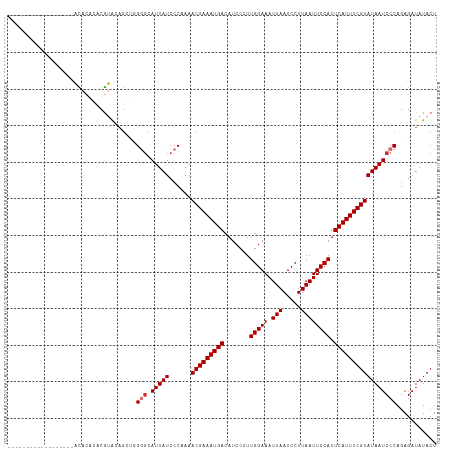
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:42:49 2011