
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 18,138,721 – 18,138,817 |
| Length | 96 |
| Max. P | 0.805406 |

| Location | 18,138,721 – 18,138,817 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 74.30 |
| Shannon entropy | 0.46419 |
| G+C content | 0.48525 |
| Mean single sequence MFE | -32.36 |
| Consensus MFE | -11.42 |
| Energy contribution | -11.62 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.805406 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 18138721 96 + 21146708 -UACUUGUCCUAGACAACUUUGCA-GUCAGC-------UGGGAACAGCUGAAACGACGCCCAG-----GCGAUGCGAGAGAUACGUAUCUGCUGGAUGC---AAACUGUAUCU -...(((((...))))).((((((-.(((((-------((....))))))).....(((....-----))).))))))(((((((....(((.....))---)...))))))) ( -33.10, z-score = -2.83, R) >droSim1.chr2R 16738348 96 + 19596830 -UACUUGUCCUAGACAACUUUGCA-GUCAGC-------UGGGAACAGCUGAAACGACGCCCAG-----GCGAUGCGAGAGAUACGUAUCUGCUGGAUGC---AAACUGUAUCU -...(((((...))))).((((((-.(((((-------((....))))))).....(((....-----))).))))))(((((((....(((.....))---)...))))))) ( -33.10, z-score = -2.83, R) >droSec1.super_9 1449084 96 + 3197100 -UACUUGUCCUAGACAACUUUGCA-GUCAGC-------UGGGAACAGCUGAAACGACGCCCAG-----GCGAUGCGAGAGAUACGUAUCUGCUGGAUGC---AAACUGUAUCU -...(((((...))))).((((((-.(((((-------((....))))))).....(((....-----))).))))))(((((((....(((.....))---)...))))))) ( -33.10, z-score = -2.83, R) >droYak2.chr2R 17888002 96 + 21139217 -UACUUGUCCUAGACAACAUUGCC-AUCAGC-------UGGGAACAGCUGAAACGACGGCCAG-----GCGAUGCGAGAGAUACGUAUCUGUUGGAUAC---AAACUGUAUCU -...(((((...)))))(((((((-.(((((-------((....)))))))...(.....).)-----))))))(((.((((....)))).)))(((((---.....))))). ( -33.30, z-score = -3.65, R) >droEre2.scaffold_4845 12279860 101 + 22589142 GUACUUGUCCUAGACAAGUUUGCA-GUCAGC-------UGGGAACAGCUGAAGCGAGGGCCAGCGAUGGCGAUGCGAGAGAUACGUAUCUGUUGGAUGC---AA-CUGUAUCU ..(((((((...)))))))((((.-.(((((-------((....))))))).))))..((((....))))((((((.......))))))....((((((---..-..)))))) ( -39.80, z-score = -3.05, R) >droAna3.scaffold_13266 2147594 92 + 19884421 AUGGCGGUCCUAGAAAA--------GUCAGC-------UGGGAACAGCUGUAAAAAUGCUAAG------CGAUGCGGAAGAUAUGUAUCUGCGGGAUGGUGAAAAUUGUAUCU .(.((.(((((......--------..((((-------((....))))))............(------(((((((.......)))))).))))))).)).)........... ( -27.70, z-score = -2.78, R) >dp4.chr3 4778754 104 - 19779522 AUGGAGGUCCUAGACAAGUCAGCAGGUCUACGUCGAAGUCGGAACAGCUGAAAUGCUGAUGAG------AGAUGCAAAAGAUAUGUAUCUAUGGGAUGG-GUGUACUGCGU-- (((.((..((((..(..(((((((..((..(.(((....)))....)..))..)))))))...------(((((((.......)))))))...)..)))-)....)).)))-- ( -29.40, z-score = -1.18, R) >droPer1.super_4 4050403 104 + 7162766 AUGGAGGUCCUAGACAAGUCAGCAGGUCUACGUCGAAGUCGGAACAGCUGAAAUGCUGAUGAG------AGAUGCAAAAGAUAUGUAUCUAUGGGAUGG-GUGUACUGCGU-- (((.((..((((..(..(((((((..((..(.(((....)))....)..))..)))))))...------(((((((.......)))))))...)..)))-)....)).)))-- ( -29.40, z-score = -1.18, R) >consensus _UACUUGUCCUAGACAACUUUGCA_GUCAGC_______UGGGAACAGCUGAAACGACGCCCAG_____GCGAUGCGAGAGAUACGUAUCUGCUGGAUGC___AAACUGUAUCU ......((((................(((((...............)))))...................((((((.......))))))....))))................ (-11.42 = -11.62 + 0.20)
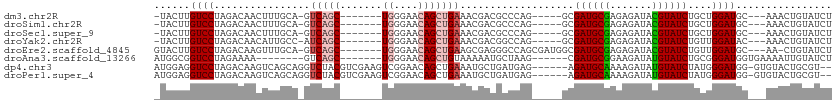

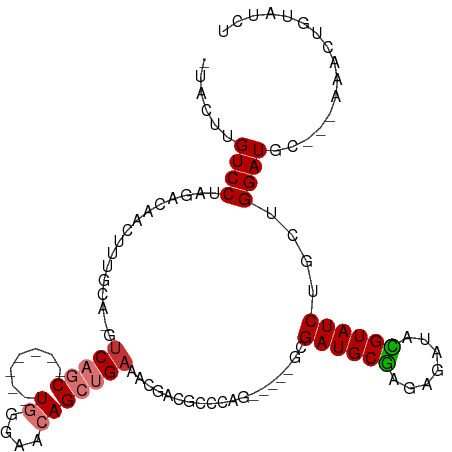

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:42:44 2011