
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 17,966,995 – 17,967,109 |
| Length | 114 |
| Max. P | 0.830149 |

| Location | 17,966,995 – 17,967,109 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 137 |
| Reading direction | reverse |
| Mean pairwise identity | 78.15 |
| Shannon entropy | 0.37509 |
| G+C content | 0.45453 |
| Mean single sequence MFE | -29.20 |
| Consensus MFE | -13.11 |
| Energy contribution | -14.44 |
| Covariance contribution | 1.32 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.830149 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 17966995 114 - 21146708 UUGGCACUUCUCACGUGCGCACCAU----GCUCCUCCA-AUUCGAUUAGAUAGCCCC------AUUGCGGACAUU--UUGAGUAAUCUCAAUGGUCGCAAC----------AAAACCUCUAAUUGUAAAUCACAGAC (((((((.......))))((.....----)).....))-)..((((((((.......------.(((((..((((--..((....))..))))..))))).----------......))))))))............ ( -25.06, z-score = -1.50, R) >droSim1.chr2R 16597041 112 - 19596830 --GGCACUUCUCACGUGCGCACCAU----GCUCCUCCA-AGUCGAUUAGAUAGCCCC------AUUGCGGCCAUU--UUGAGUAAUCUCAAUGGUCGCAAC----------AAAACCUCUAAUUGUAAAUCACAGAC --.((((.......))))((.....----)).......-...((((((((.......------.(((((((((((--..((....))..))))))))))).----------......))))))))............ ( -30.76, z-score = -3.35, R) >droSec1.super_9 1278861 112 - 3197100 --GGCACUUCUCACGUGCACACCAU----GCUCCUCCA-AGUCGAUUAGAUAGCCCC------AUUGCGGCCAUU--UUGAGUAAUCUCAAUGGUCGCAAC----------AAAACCUCUAAUUGUAAAUCACAGAC --.((((.......)))).......----.........-...((((((((.......------.(((((((((((--..((....))..))))))))))).----------......))))))))............ ( -29.66, z-score = -3.38, R) >droYak2.chr2R 9927635 111 + 21139217 --GGCAAUUCUCACGUGCCAACCAU----GCUCUUCCA-AGUCGAUUAGGUAG-CCC------AAUGCGGCCAUU--UUGAGUAAUCUCAAUGGUCGCAAC----------AAAACCUCUAAUUGUAAAUCACAGAC --((((.........))))......----..(((....-...((((((((...-...------..((((((((((--..((....))..))))))))))..----------......))))))))........))). ( -28.25, z-score = -2.61, R) >droEre2.scaffold_4845 12105542 103 - 22589142 -----GGCUCUC-----CUCGCCAU----GCUCGUACA-AGUCGAUUAGAUAG-CCC------AUUGCGGCCAUU--UUGAGUAAUCUCAAUGUACGCAAC----------AAAACCUCUAAUUGUAAAUCACAGAC -----(((....-----...)))..----....((...-...((((((((...-...------.(((((..((((--..((....))..))))..))))).----------......))))))))......)).... ( -23.54, z-score = -2.08, R) >droAna3.scaffold_13266 11332712 137 - 19884421 UGGGCGCUUCUCACGUGCCCCCCAUUUAAACUCCUCCAUGAUUGAUUAGAUAGCCGCAGGUUAAUUGCCGACAUUGGGGGAGUACGUUCCAUAUACAUAAUCUGAAAACAGAAAAACCUAAAUGGCAAAUCACUGAC .((((((.......)))))).(((((((.(((((((((((.(((..(((.(((((...))))).))).))))).))))))))).................((((....))))......)))))))............ ( -37.90, z-score = -2.17, R) >consensus __GGCACUUCUCACGUGCCCACCAU____GCUCCUCCA_AGUCGAUUAGAUAGCCCC______AUUGCGGCCAUU__UUGAGUAAUCUCAAUGGUCGCAAC__________AAAACCUCUAAUUGUAAAUCACAGAC ...((((.......))))........................(((((((((.............((((((((((....((((....)))))))))))))).............)...))))))))............ (-13.11 = -14.44 + 1.32)
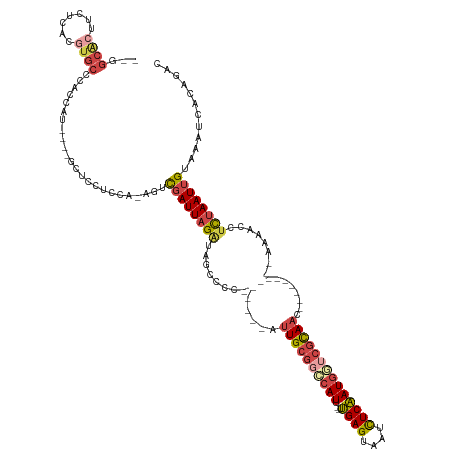


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:42:27 2011