
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 17,945,945 – 17,946,014 |
| Length | 69 |
| Max. P | 0.865524 |

| Location | 17,945,945 – 17,946,014 |
|---|---|
| Length | 69 |
| Sequences | 12 |
| Columns | 81 |
| Reading direction | reverse |
| Mean pairwise identity | 79.12 |
| Shannon entropy | 0.42317 |
| G+C content | 0.32370 |
| Mean single sequence MFE | -12.90 |
| Consensus MFE | -7.20 |
| Energy contribution | -7.18 |
| Covariance contribution | -0.01 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.865524 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 17945945 69 - 21146708 --------UUUUUUCCGUAUUAUU--UUUCUGCGGUGACAAACUUGGCAACGAGUCGC--AGUUCCUUUUUAAUUUAUUUG --------................--...(((((....)..(((((....))))).))--))................... ( -13.70, z-score = -2.33, R) >droPer1.super_4 3630307 73 + 7162766 -----UUUUUUCCUUAUUAUUAUU--UUUCUGCGGUGACAAACUUGGCAACGAGUUGC-GACUUCUUUUUUAAUUUAUUUG -----...................--....((((....).((((((....))))))))-)..................... ( -11.80, z-score = -2.32, R) >dp4.chr3 15513970 73 - 19779522 -----UUUUUUCCUUAUUAUUAUU--UUUCUGCGGUGACAAACUUGGCAACGAGUUGC-GACUUCUUUUUUAAUUUAUUUG -----...................--....((((....).((((((....))))))))-)..................... ( -11.80, z-score = -2.32, R) >droAna3.scaffold_13266 11313973 71 - 19884421 --------UUUUUUCGAUAUUAUU--UUUCUGCGGCGACAAACUUGGCAGCCGGUUGCAUUCUUAUUUUUCCAUUUAUUUU --------................--......((((..((....))...))))............................ ( -7.30, z-score = 0.50, R) >droEre2.scaffold_4845 12084159 69 - 22589142 --------UUUUUUCCGUAUUAUU--UUUCUGCGGUGACAAACUUGGCAACGAGUCGC--AGUUCCUUUUUAAUUUAUUUG --------................--...(((((....)..(((((....))))).))--))................... ( -13.70, z-score = -2.33, R) >droYak2.chr2R 9906186 69 + 21139217 --------UUUUUUCCGUAUUAUU--UUUCUGCGGUGACAAACUUGGCAACGAGUCGC--AGUUCCUUUUUAAUUUAUUUG --------................--...(((((....)..(((((....))))).))--))................... ( -13.70, z-score = -2.33, R) >droSec1.super_9 1257842 68 - 3197100 --------UUUUUUCCGUAUUAUU--UUUCUGCGGUGACAAACUUGGCAACGAGUCGC--AGUU-CUUUUUAAUUUAUUUG --------................--...(((((....)..(((((....))))).))--))..-................ ( -13.70, z-score = -2.41, R) >droSim1.chr2R 16575712 69 - 19596830 --------UUUUUUCCGUAUUAUU--UUUCUGCGGUGACAAACUUGGCAACGAGUCGC--AGUUCCUUUUUAAUUUAUUUG --------................--...(((((....)..(((((....))))).))--))................... ( -13.70, z-score = -2.33, R) >droWil1.scaffold_180701 550624 76 + 3904529 UUUCCUUCAUAUUUUCAUUUUGUUGCUGUCUGCGGUGACAAACUUGGCAACGAGUUGC-AA----UUUUUUAAUUUAUUUG ....................(((..(((....)))..)))((((((....))))))..-..----................ ( -16.50, z-score = -2.55, R) >droVir3.scaffold_12875 11579130 65 - 20611582 --------UUUUUUGCGUUUCAUU--UUUCUGUGGUGACAAACUUGGCAACGAGUUGCA-----AUUUUUAAAUUUAUUU- --------....(((((((((((.--.....)))).))).((((((....)))))))))-----)...............- ( -14.10, z-score = -2.22, R) >droMoj3.scaffold_6496 11567910 69 + 26866924 --------UUGUUGUUGUUAUAUU-UGUUAUGUGGUGACAAACUUGGCAACGAGUUGCAAA---UUUUUUCAACUUAUUUU --------.....((((....(((-(((....((....))((((((....)))))))))))---).....))))....... ( -17.00, z-score = -2.65, R) >droGri2.scaffold_15245 11682656 66 + 18325388 --------GCUUUUCCAUUUUAUU--UUUCUGUGGUGACAAACUUGGCAAAGAGCUGCA-----AUUUUUAAAUUUAUUUG --------(((((((((.......--......((....))....)))..))))))....-----................. ( -7.83, z-score = 0.56, R) >consensus ________UUUUUUCCGUAUUAUU__UUUCUGCGGUGACAAACUUGGCAACGAGUUGC__AGUUCUUUUUUAAUUUAUUUG ...............................(((....)..(((((....))))).))....................... ( -7.20 = -7.18 + -0.01)
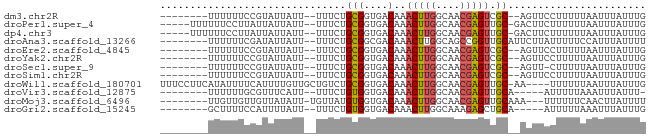

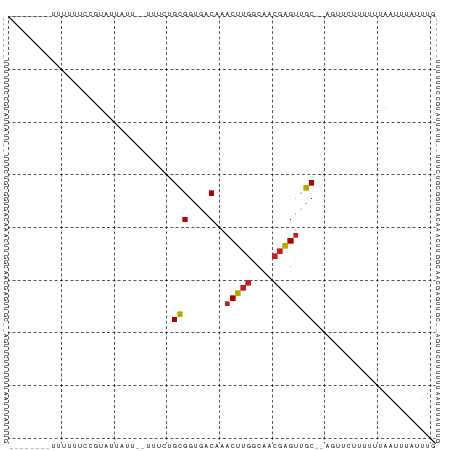
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:42:26 2011