
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 17,907,859 – 17,907,972 |
| Length | 113 |
| Max. P | 0.559251 |

| Location | 17,907,859 – 17,907,972 |
|---|---|
| Length | 113 |
| Sequences | 7 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 74.07 |
| Shannon entropy | 0.53429 |
| G+C content | 0.40845 |
| Mean single sequence MFE | -20.86 |
| Consensus MFE | -9.71 |
| Energy contribution | -11.63 |
| Covariance contribution | 1.92 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.559251 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 17907859 113 + 21146708 GACAUUAUGCUUAAUUGCAGCGUCAGUUUGAAUUUAACUGAGAUUGCCGCUCCUCAGCCUCUGCAUCAUU-CCAUUUUC-UUUUUGCCUCCAUUGUUUCUGCGGUGGAUUUUUUC ......((((......((((..((((((.......))))))..)))).(((....)))....))))....-........-........((((((((....))))))))....... ( -22.80, z-score = -0.72, R) >droSim1.chr2R 16549464 107 + 19596830 GACAUUAUGCUUAAUUGCAGCGUCAGUUUGAAUUUAACUGAGAUUGCCGCUCCUCAGCCCCA-------UUCCAUUUUC-UUUUUGCCUCCAUUGUUUCUGCGGUGGAUUUUUCC .......(((......)))(((((((((.......))))))((.((..(((....)))..))-------.)).......-....))).((((((((....))))))))....... ( -21.30, z-score = -1.07, R) >droSec1.super_9 1219845 113 + 3197100 GACAUUAUGCUUAAUUGCAGCGUCAGUUUGAAUUUAACUGAGAUUGCCGCUCCUCAGCCUCAUCCCCAUU-CCAUUUUC-CUUUUGCCUCCAUUGUUUCUGCGGUGGAUUUUUCC ........(((.....((.(((((((((.......))))))...))).)).....)))............-........-........((((((((....))))))))....... ( -21.00, z-score = -0.82, R) >droYak2.chr2R 9866700 115 - 21139217 GACAUUAUGCUUAAUUGCAGCGUCAGUUUGAAUUUAACUGAGAUUGCCGCUCCUCAGCCUCAGUUUCAGUCUCAUUUUCCUUUUUACCUCCAUUGUUUCUGCGGUGGAUUUUUCC (((....(((......)))..)))............((((((((((..(((....)))..))))))))))..................((((((((....))))))))....... ( -28.10, z-score = -2.89, R) >droEre2.scaffold_4845 12042876 113 + 22589142 GACAUUAUGCUUAAUUGCAGCGUCAGUUUGAAUUUAACUGAGAUUGCCGCUCCUCAGCCUCAGCUUCAGU-UCAUUUUC-UUUUUGCCUCCAUUGUUUCUGCGGUGGAUUUUUCC (((....(((......)))..))).....(((..((((((((.(((..(((....)))..))).))))))-).)..)))-........((((((((....))))))))....... ( -26.10, z-score = -1.44, R) >droPer1.super_323 20460 102 + 21222 GACAUUAUGCUUAAUUGCAGCGUCUGUUUGAAUUUAACCGAGAUUCAAGCCUUACC-CUUAUCCCAUGGCCUCUUUUUACCCACCAUGUAUACAUACUUAUCC------------ (((....(((......)))..))).((((((((((.....))))))))))......-.......(((((..............)))))...............------------ ( -17.24, z-score = -2.58, R) >droWil1.scaffold_180701 3294865 101 + 3904529 GACAUUAUACUUAAUUGCAGCAUCAGUUUGAAUUCAACCGAGAUU----CUCUACAACAUCCUCU----CAUCCUCUUAAUUUUUUUUUUUAGUCGUUUU----UGGGUUUUC-- ........((((((..((.((....(.(((....))).)((((..----.............)))----)......................)).))..)----)))))....-- ( -9.46, z-score = 0.49, R) >consensus GACAUUAUGCUUAAUUGCAGCGUCAGUUUGAAUUUAACUGAGAUUGCCGCUCCUCAGCCUCAGCUUCAGU_UCAUUUUC_UUUUUGCCUCCAUUGUUUCUGCGGUGGAUUUUUCC ................((((..((((((.......))))))..)))).........................................((((((((....))))))))....... ( -9.71 = -11.63 + 1.92)

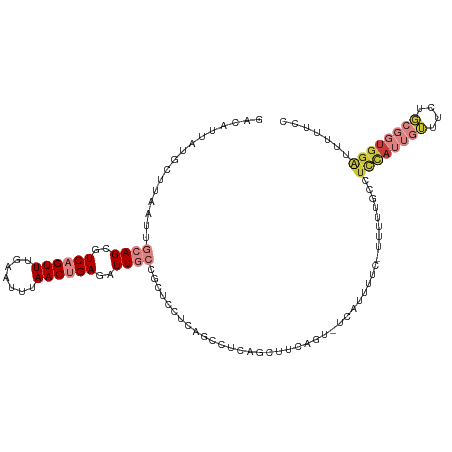
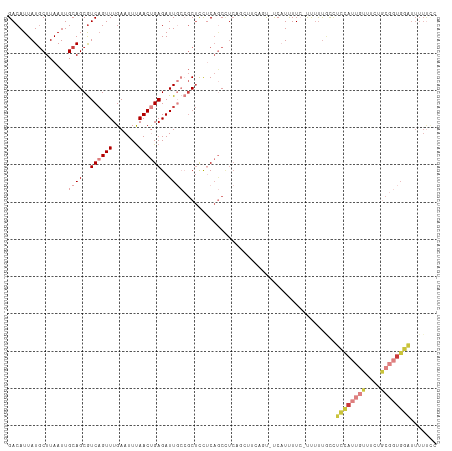
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:42:16 2011