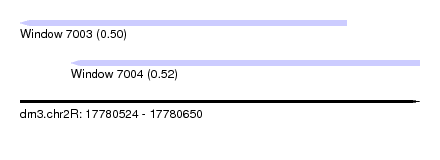

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 17,780,524 – 17,780,650 |
| Length | 126 |
| Max. P | 0.523515 |

| Location | 17,780,524 – 17,780,627 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 78.75 |
| Shannon entropy | 0.38444 |
| G+C content | 0.32729 |
| Mean single sequence MFE | -18.61 |
| Consensus MFE | -12.11 |
| Energy contribution | -14.03 |
| Covariance contribution | 1.92 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr2R 17780524 103 - 21146708 GGGUCGAGUGCUUUUCAGUUUUCGAUCAAUUUCAAUUUGGCUUAGCCAAUUAGUUUGUAUAAAUAUUUAAGCUUUUUGCAGCUCAAUUUAGGUCCUCUUUUUA----------- .(((((((.(((....))).))))))).((((.((((.((((..((.((..((((((..........))))))..)))))))).)))).))))..........----------- ( -19.50, z-score = -1.23, R) >droSim1.chr2R 16415988 103 - 19596830 GGGUCGAGUGCUUUUCAGUUUUCGAUCAAUUUCAAUUUGGCUUAGCUAAUUAGUUUGUAUAAAUAUUUAAGCUUUUGGCAGUUCAAUUUAGGUCAUCUUUUUA----------- .(((((((.(((....))).)))))))..........((((((((((((..((((((..........)))))).))))).........)))))))........----------- ( -21.30, z-score = -2.53, R) >droSec1.super_9 1095597 103 - 3197100 GGGUCGAGUGCUUUUCAGUUUUCGAUCAAUUUCAAUUUGGCUUAGCUAAUUAGUUUGUAUAAAUAUUUAAGCUUUUGGCAGUUCAAUUUACGUCCUCUUUUUA----------- .(((((((.(((....))).)))))))......((((..(((..(((((..((((((..........)))))).))))))))..))))...............----------- ( -19.30, z-score = -1.63, R) >droYak2.chr2R 9734912 103 + 21139217 GGGUCGAGUGGCUUUCAAUCUUCGAUCAAUUUCAAUUUGGCUCAGCUAAUUAGUUUGUAUAAAUAUUUAAGCUUUUGGCAGCUCAAUUUAGGUCCUUCUUUUA----------- ((..(...(((((..........((((((.......)))).)).(((((..((((((..........)))))).)))))))).)).....)..))........----------- ( -19.20, z-score = -0.70, R) >droEre2.scaffold_4845 11914308 103 - 22589142 GGGUCGAGUGCCCCUCAGUCUUCGAUCAAUUUCAAUUUGGCUUAGCUAAUUAGUUUGGAUAAAUAUUUAAGCUUUUGGCAGCUCAAUUUACGUCCUCUUUUUG----------- .(((((((.((......)).)))))))......((((.((((..(((((..(((((((((....))))))))).))))))))).))))...............----------- ( -23.80, z-score = -1.69, R) >droAna3.scaffold_13266 11159715 102 - 19884421 ------------UUUCAAUUUCCUCUAAUUUCUAAUUAUUCAAAAUUAAUUAGUUUGUCCAAAUAUUUAAACUUUUGACAGCCCAAUUUUGUUUUUUGCUCUGGUCCUGAUUUC ------------..(((..............(((((((........)))))))...(.(((.......((((..(((......)))....)))).......))).).))).... ( -8.54, z-score = 0.11, R) >consensus GGGUCGAGUGCUUUUCAGUUUUCGAUCAAUUUCAAUUUGGCUUAGCUAAUUAGUUUGUAUAAAUAUUUAAGCUUUUGGCAGCUCAAUUUAGGUCCUCUUUUUA___________ .(((((((.((......)).)))))))......((((.((((..(((((..((((((..........)))))).))))))))).)))).......................... (-12.11 = -14.03 + 1.92)
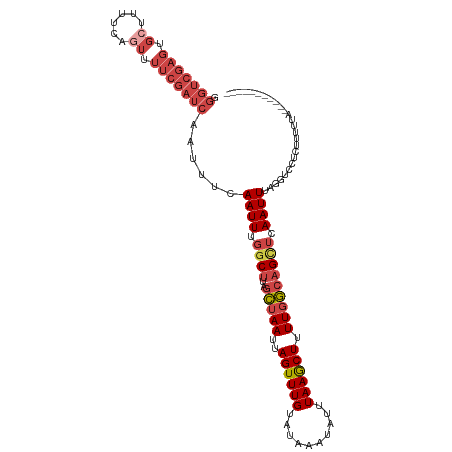


| Location | 17,780,540 – 17,780,650 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 92.73 |
| Shannon entropy | 0.12515 |
| G+C content | 0.32614 |
| Mean single sequence MFE | -19.67 |
| Consensus MFE | -17.84 |
| Energy contribution | -18.00 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.523515 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 17780540 110 - 21146708 UACAUUUUUCAAAUCCUUUCAAAGGGUCGAGUGCUUUUCAGUUUUCGAUCAAUUUCAAUUUGGCUUAGCCAAUUAGUUUGUAUAAAUAUUUAAGCUUUUUGCAGCUCAAU ...((((..(((((..........(((((((.(((....))).))))))).........(((((...)))))...)))))...)))).....((((......)))).... ( -18.70, z-score = -1.18, R) >droSim1.chr2R 16416004 110 - 19596830 UACAUUUUUCAAAUCCUUUCAAAGGGUCGAGUGCUUUUCAGUUUUCGAUCAAUUUCAAUUUGGCUUAGCUAAUUAGUUUGUAUAAAUAUUUAAGCUUUUGGCAGUUCAAU ........((((((..........(((((((.(((....))).))))))).......))))))....(((((..((((((..........)))))).)))))........ ( -18.93, z-score = -1.34, R) >droSec1.super_9 1095613 110 - 3197100 UACAUUUUUCAAAACCUUUCAAAGGGUCGAGUGCUUUUCAGUUUUCGAUCAAUUUCAAUUUGGCUUAGCUAAUUAGUUUGUAUAAAUAUUUAAGCUUUUGGCAGUUCAAU ........................(((((((.(((....))).))))))).........((((((..(((((..((((((..........)))))).))))))).)))). ( -18.50, z-score = -1.00, R) >droYak2.chr2R 9734928 110 + 21139217 UACAUUUUUCAUAUCCCUUCACAGGGUCGAGUGGCUUUCAAUCUUCGAUCAAUUUCAAUUUGGCUCAGCUAAUUAGUUUGUAUAAAUAUUUAAGCUUUUGGCAGCUCAAU ..............((((....))))..((((..............((((((.......)))).)).(((((..((((((..........)))))).))))).))))... ( -18.80, z-score = -0.85, R) >droEre2.scaffold_4845 11914324 109 - 22589142 UACAUUUUUCAUAU-CCUCCACAGGGUCGAGUGCCCCUCAGUCUUCGAUCAAUUUCAAUUUGGCUUAGCUAAUUAGUUUGGAUAAAUAUUUAAGCUUUUGGCAGCUCAAU ..............-.........(((((((.((......)).))))))).........((((((..(((((..(((((((((....))))))))).)))))))).))). ( -23.40, z-score = -1.57, R) >consensus UACAUUUUUCAAAUCCUUUCAAAGGGUCGAGUGCUUUUCAGUUUUCGAUCAAUUUCAAUUUGGCUUAGCUAAUUAGUUUGUAUAAAUAUUUAAGCUUUUGGCAGCUCAAU ........................(((((((.((......)).))))))).........((((((..(((((..((((((..........)))))).)))))))).))). (-17.84 = -18.00 + 0.16)
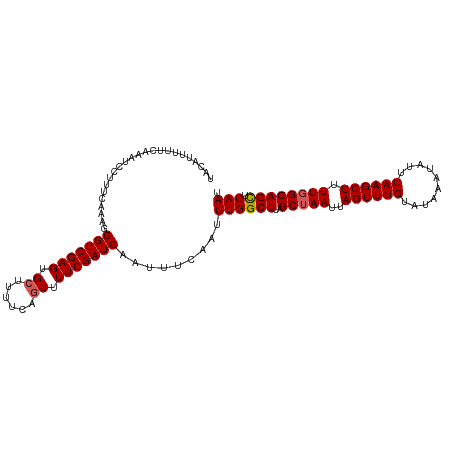



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:41:54 2011