
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 17,757,956 – 17,758,046 |
| Length | 90 |
| Max. P | 0.745024 |

| Location | 17,757,956 – 17,758,046 |
|---|---|
| Length | 90 |
| Sequences | 11 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 55.50 |
| Shannon entropy | 0.94926 |
| G+C content | 0.53888 |
| Mean single sequence MFE | -21.30 |
| Consensus MFE | -8.07 |
| Energy contribution | -7.99 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -0.64 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.745024 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 17757956 90 + 21146708 ----CCCUCGAGAGGACUACCCCCCUCACCACCCUCCAAAAGUCUUCGUCUUC---GGAG----CGUGAGGCGUGCAUAAGUGAAAUGUGCACAAGAUACA ----.((((..((((........))))...((.((((..(((.......))).---))))----.)))))).(((((((.......)))))))........ ( -25.30, z-score = -1.80, R) >droSec1.super_9 1073078 79 + 3197100 ----CCCUCGAGAGGACCACCCCCC-----------CGAAAGUCACCGUCUUC---GGAG----CGUGAGGCGUGCAUAAGUGAAAUGUGCACAAGAUACA ----.(((((.((((((........-----------(....).....))))))---....----..))))).(((((((.......)))))))........ ( -20.32, z-score = -0.86, R) >droYak2.chr2R 9711205 78 - 21139217 ----CCCUCGAGAGGACCACCCCCC------------GAAAGCCAACAAGCUG---GGAG----CGUGAGGCGUGCAUAAGUGAAAUGUGCACAAGAUACA ----.(((((...((.....))(((------------...(((......))))---))..----..))))).(((((((.......)))))))........ ( -20.00, z-score = -0.92, R) >droEre2.scaffold_4845 11891448 78 + 22589142 ----CCCUCGAGGGGACCAGCCCCC------------GAAAGCCGCCGUCUUC---GGAG----CGUGAGGCGUGCAUAAGUGAAAUGUGCACAAGAUACA ----...(((.((((.....)))))------------))..(((.(((((...---.)).----)).).)))(((((((.......)))))))........ ( -29.50, z-score = -2.43, R) >droAna3.scaffold_13266 11139239 88 + 19884421 ------CCCUCCAACACCAACUCCAACUCGGAGUGCCAGUCCCUUCUGCCAGC---UAAG----CGUGAGGCGUGCAUAAGUGAAAUGUGCACAAGAUACA ------.(((((.......(((((.....)))))(((((......)))...))---....----.).)))).(((((((.......)))))))........ ( -22.10, z-score = -0.84, R) >dp4.chr3 15311152 80 + 19779522 ----CCCUCUCUGCCGCCGUCUCAGACUCG-----CCAG--CGAUCCACCA------AAG----CGUGAGGCGUGCAUAAGUGAAAUGUGCACAAGAUACC ----.....(((...(((......((.(((-----....--)))))(((..------...----.))).)))(((((((.......))))))).))).... ( -18.70, z-score = -0.17, R) >droPer1.super_4 3422508 80 - 7162766 ----CCCUCUCUGCCGCCGUCUCAGACUCG-----CCAG--CGAUCCACCA------AAG----CGUGAGGCGUGCAUAAGUGAAAUGUGCACAAGAUACC ----.....(((...(((......((.(((-----....--)))))(((..------...----.))).)))(((((((.......))))))).))).... ( -18.70, z-score = -0.17, R) >droWil1.scaffold_180701 3069204 96 + 3904529 CAUUCCAUCAGGCUGCAGACUUCACUUGCUUGGAUGCUGUUCGUCCUCCUUCUCCUUUUGCUUCUCCUCCUUGGGCAUAAGUGAAAUGUGCACAAG----- .............((((.(.((((((((...(((((.....)))))............(((((.........))))))))))))).).))))....----- ( -20.30, z-score = 0.19, R) >droGri2.scaffold_15245 3469972 97 + 18325388 UUGUUGUUUGUGGCAGCCCCUCCCCAAAUUUGGCAUUUGCUGUGCCAGCCUCG----CAGUUUCCUUGAAGCGUGCAUAAGUGAAAUGUGCGCAAGAUACU .(((.(((((.((.((...)).)))))))(((((((.....)))))))....)----))(((((...)))))(((((((.......)))))))........ ( -22.70, z-score = 0.65, R) >droMoj3.scaffold_6496 10203748 98 - 26866924 ACCUCCACCAUGCCCUCCUCUCCACAAC---AGCAAUUGCUGUAGCAGCCUUUAGUUCAGUUUCCUUCAAGCGUGCAUAAGUGAAAUGUGCGCCAGAUACU ..........................((---(((....))))).......................((..(((..(((.......)))..)))..)).... ( -16.50, z-score = 0.15, R) >droVir3.scaffold_12875 10324504 88 + 20611582 ACGUUGACUG----AACUGCCCCCAAAA----GCAUUUGCUGUGCCAGCCUC-----CAGUUUCCUUCAAGCGUGCAUAAGUGAAAUGUGCGCAAGAUACU ..(..(((((----...(((........----)))...((((...))))...-----)))))..).....(((..(((.......)))..)))........ ( -20.20, z-score = -0.82, R) >consensus ____CCCUCGAGACGACCACCCCCCAC________CCAGAAGUCUCCGCCUUC____GAG____CGUGAGGCGUGCAUAAGUGAAAUGUGCACAAGAUACA ........................................................................(((((((.......)))))))........ ( -8.07 = -7.99 + -0.08)

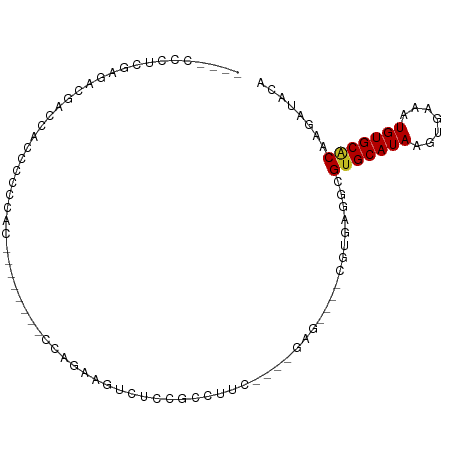
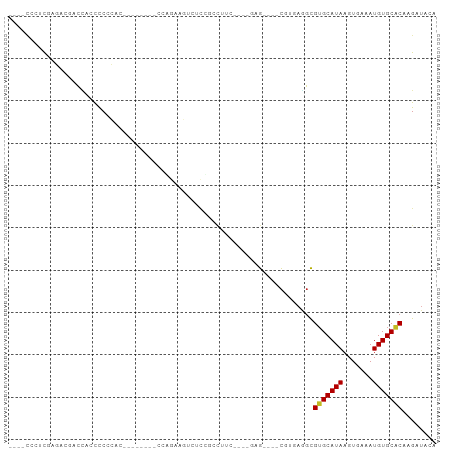
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:41:48 2011