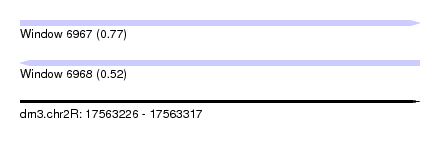

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 17,563,226 – 17,563,317 |
| Length | 91 |
| Max. P | 0.771626 |

| Location | 17,563,226 – 17,563,317 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 90.62 |
| Shannon entropy | 0.16867 |
| G+C content | 0.46941 |
| Mean single sequence MFE | -27.76 |
| Consensus MFE | -21.84 |
| Energy contribution | -21.92 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.771626 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 17563226 91 + 21146708 GUGCAUGGCAUACCAUAACGAUUCCGAUUUCAAAUCGGUUUGGUUUCGGGGCGGCUUUAUCAGCGCCUCGCACGCAGCUAUUUAUAGUUGA----- ((((((((....))))(((.(..(((((.....)))))..).)))..((((((.((.....)))))))))))).((((((....)))))).----- ( -32.20, z-score = -3.14, R) >droSim1.chr2R 16212115 90 + 19596830 GUGCAUGGCAUACCAUAACGAUUCCGAUUUCAAAUCGGUU-GGUUUUGGGGCGGCUUUAUCAGCGCCUCACACGCAGCUAUUUAUAGUUGA----- (((...(((...(((.(((.(..(((((.....))))).)-.))).)))....(((.....))))))...))).((((((....)))))).----- ( -26.80, z-score = -1.85, R) >droSec1.super_9 889076 90 + 3197100 GUGCAUGGCAUACCAUAACGAUUCCGAUUUCAAAUCGGUU-GGUUUUGGGGCGGCUUUAUCAGCGCCUCACACGCAGCUAUUUAUAGUUGA----- (((...(((...(((.(((.(..(((((.....))))).)-.))).)))....(((.....))))))...))).((((((....)))))).----- ( -26.80, z-score = -1.85, R) >droYak2.chr2R 9530945 94 - 21139217 GUGCAUGGCAUACCAUAACGAUUCCGAUUUCAAAUCGCUU--GCUUUGGGGCGGGUUUAUCAGCGCCUCACACGCAGCUAUUUAUAGUUGGGAGUG ((..((((....)))).))..((((((((..((((.(((.--((..(((((((.(.....)..)))))))...))))).))))..))))))))... ( -29.20, z-score = -1.75, R) >droEre2.scaffold_4845 11711563 93 + 22589142 GUGCAUGGCAUACCAUA-CAAUUCCGAUUUCAAAUCGGUU--GGUUUGGGGCGGGUUUAUCAGCGCCUCACACGCAGCUAUUUAUAGUUACAUAUA ..((((((....)))).-(((..(((((.....)))))))--)((.(((((((.(.....)..))))))).))))(((((....)))))....... ( -23.80, z-score = -1.45, R) >consensus GUGCAUGGCAUACCAUAACGAUUCCGAUUUCAAAUCGGUU_GGUUUUGGGGCGGCUUUAUCAGCGCCUCACACGCAGCUAUUUAUAGUUGA_____ (((.((((....)))).......(((((.....)))))........(((((((..........)))))))))).((((((....))))))...... (-21.84 = -21.92 + 0.08)
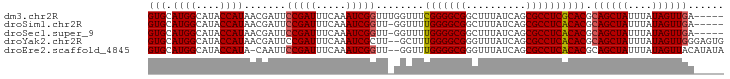


| Location | 17,563,226 – 17,563,317 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 90.62 |
| Shannon entropy | 0.16867 |
| G+C content | 0.46941 |
| Mean single sequence MFE | -23.64 |
| Consensus MFE | -19.34 |
| Energy contribution | -19.86 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.524499 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 17563226 91 - 21146708 -----UCAACUAUAAAUAGCUGCGUGCGAGGCGCUGAUAAAGCCGCCCCGAAACCAAACCGAUUUGAAAUCGGAAUCGUUAUGGUAUGCCAUGCAC -----...............((((((((.((((((.....)).)))).))..((((...((((((.......))))))...))))....)))))). ( -24.50, z-score = -1.62, R) >droSim1.chr2R 16212115 90 - 19596830 -----UCAACUAUAAAUAGCUGCGUGUGAGGCGCUGAUAAAGCCGCCCCAAAACC-AACCGAUUUGAAAUCGGAAUCGUUAUGGUAUGCCAUGCAC -----...............((((((...((((((.....)).))))(((.(((.-..(((((.....)))))....))).))).....)))))). ( -23.30, z-score = -1.59, R) >droSec1.super_9 889076 90 - 3197100 -----UCAACUAUAAAUAGCUGCGUGUGAGGCGCUGAUAAAGCCGCCCCAAAACC-AACCGAUUUGAAAUCGGAAUCGUUAUGGUAUGCCAUGCAC -----...............((((((...((((((.....)).))))(((.(((.-..(((((.....)))))....))).))).....)))))). ( -23.30, z-score = -1.59, R) >droYak2.chr2R 9530945 94 + 21139217 CACUCCCAACUAUAAAUAGCUGCGUGUGAGGCGCUGAUAAACCCGCCCCAAAGC--AAGCGAUUUGAAAUCGGAAUCGUUAUGGUAUGCCAUGCAC ...(((......(((((.(((((...((.((((..........)))).))..))--.))).))))).....)))...((.((((....)))))).. ( -21.40, z-score = -0.28, R) >droEre2.scaffold_4845 11711563 93 - 22589142 UAUAUGUAACUAUAAAUAGCUGCGUGUGAGGCGCUGAUAAACCCGCCCCAAACC--AACCGAUUUGAAAUCGGAAUUG-UAUGGUAUGCCAUGCAC ((((((((.(((....))).)))))))).((((..........)))).......--..(((((.....)))))...((-(((((....))))))). ( -25.70, z-score = -2.68, R) >consensus _____UCAACUAUAAAUAGCUGCGUGUGAGGCGCUGAUAAAGCCGCCCCAAAACC_AACCGAUUUGAAAUCGGAAUCGUUAUGGUAUGCCAUGCAC ....................((((((...((((((.....)).))))(((.(((....(((((.....)))))....))).))).....)))))). (-19.34 = -19.86 + 0.52)

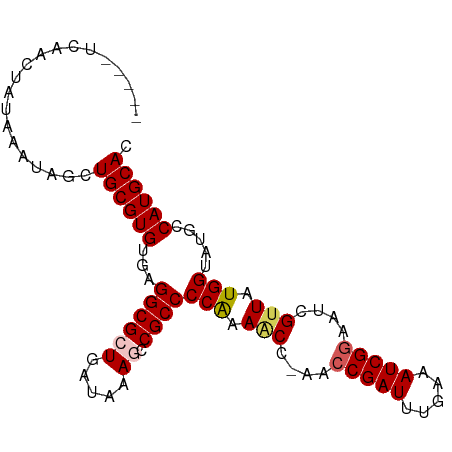
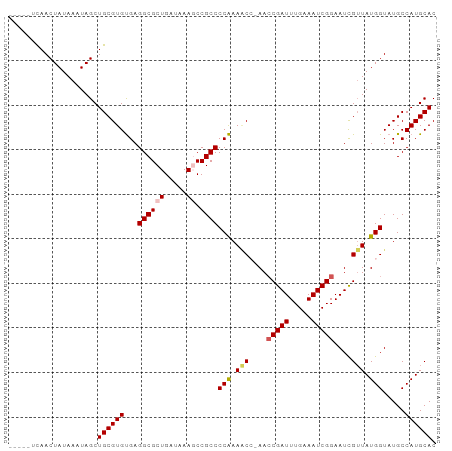
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:41:24 2011