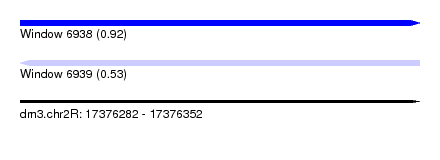

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 17,376,282 – 17,376,352 |
| Length | 70 |
| Max. P | 0.916497 |

| Location | 17,376,282 – 17,376,352 |
|---|---|
| Length | 70 |
| Sequences | 5 |
| Columns | 72 |
| Reading direction | forward |
| Mean pairwise identity | 92.98 |
| Shannon entropy | 0.11756 |
| G+C content | 0.38261 |
| Mean single sequence MFE | -19.60 |
| Consensus MFE | -15.54 |
| Energy contribution | -15.66 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.916497 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
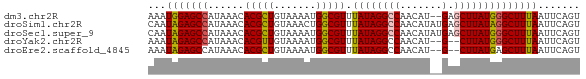
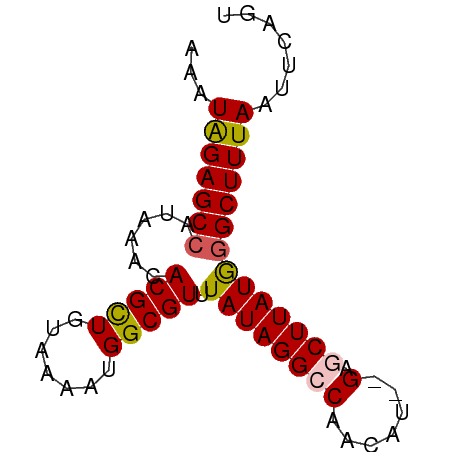
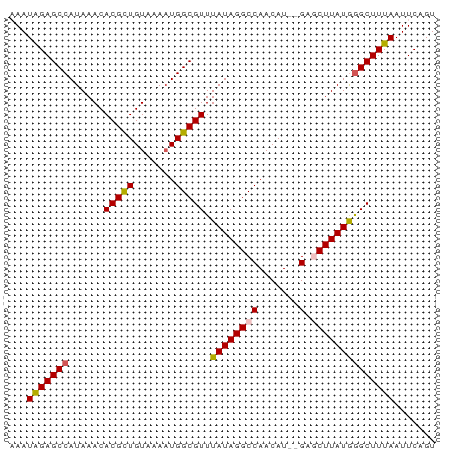
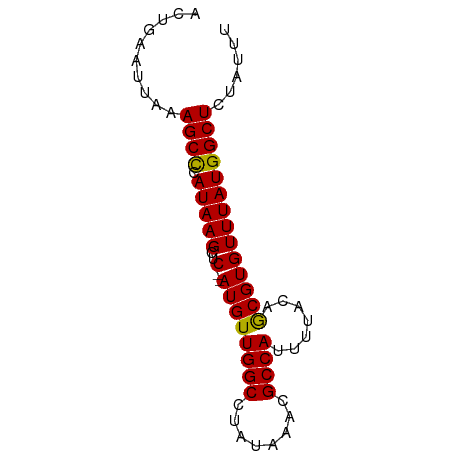
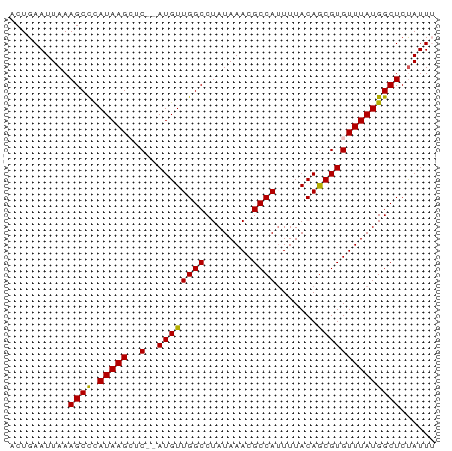
>dm3.chr2R 17376282 70 + 21146708 AAAUGGAGCCAUAAACACGCUGUAAAAUGGCGUUUAUAGGCCAACAU--GAGCUUAUGGGCUUUAAUUCAGU ...(((((((......(((((((...))))))).((((((((.....--).))))))))))))))....... ( -19.90, z-score = -2.14, R) >droSim1.chr2R 16035901 72 + 19596830 CAAUAGAGCCAUAAACACGCUGUAAACUGGCGUUUAUAGGCCAACAUAUGAGCUUAUAGGCUUUAAUUCAGU ...(((((((......(((((.......))))).(((((((((.....)).))))))))))))))....... ( -21.30, z-score = -3.29, R) >droSec1.super_9 706103 72 + 3197100 CAAUAGAGCCAUAAACACGCUGUAAAAUGGCGUUUAUAGGCCAACAUAUGAGCUUAUGGGCUUUAAUUCAGU ...(((((((......(((((((...))))))).(((((((((.....)).))))))))))))))....... ( -20.90, z-score = -2.87, R) >droYak2.chr2R 9335346 68 - 21139217 AAAUAGAGCCAUAAACACGUUGUAAAAUGGCGUUUAUAGGCCAACAU--G--CUUAUGGGCUUUAAUUCAGU ...(((((((......((((..(...)..)))).(((((((......--)--)))))))))))))....... ( -18.40, z-score = -2.35, R) >droEre2.scaffold_4845 11521073 68 + 22589142 AAAUAGAGCCAUAAACACGCUGUAAAAUGGCGUUUAUAGGCCAACAU--G--CUUAUGAGCUUUAAUUCAGU .....(((((((..((.....))...)))))((((((((((......--)--)))))))))......))... ( -17.50, z-score = -2.20, R) >consensus AAAUAGAGCCAUAAACACGCUGUAAAAUGGCGUUUAUAGGCCAACAU__GAGCUUAUGGGCUUUAAUUCAGU ...(((((((......(((((.......))))).(((((((..........))))))))))))))....... (-15.54 = -15.66 + 0.12)
| Location | 17,376,282 – 17,376,352 |
|---|---|
| Length | 70 |
| Sequences | 5 |
| Columns | 72 |
| Reading direction | reverse |
| Mean pairwise identity | 92.98 |
| Shannon entropy | 0.11756 |
| G+C content | 0.38261 |
| Mean single sequence MFE | -17.16 |
| Consensus MFE | -13.12 |
| Energy contribution | -13.36 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.533681 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
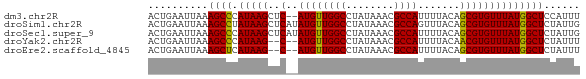
>dm3.chr2R 17376282 70 - 21146708 ACUGAAUUAAAGCCCAUAAGCUC--AUGUUGGCCUAUAAACGCCAUUUUACAGCGUGUUUAUGGCUCCAUUU ..........((((.((((((.(--.(((((((........))))....)))..).))))))))))...... ( -16.10, z-score = -1.64, R) >droSim1.chr2R 16035901 72 - 19596830 ACUGAAUUAAAGCCUAUAAGCUCAUAUGUUGGCCUAUAAACGCCAGUUUACAGCGUGUUUAUGGCUCUAUUG ..........((((.((((((.(...(((((((........))))....)))..).))))))))))...... ( -15.60, z-score = -0.89, R) >droSec1.super_9 706103 72 - 3197100 ACUGAAUUAAAGCCCAUAAGCUCAUAUGUUGGCCUAUAAACGCCAUUUUACAGCGUGUUUAUGGCUCUAUUG ..........((((.((((((.(...(((((((........))))....)))..).))))))))))...... ( -15.60, z-score = -1.34, R) >droYak2.chr2R 9335346 68 + 21139217 ACUGAAUUAAAGCCCAUAAG--C--AUGUUGGCCUAUAAACGCCAUUUUACAACGUGUUUAUGGCUCUAUUU ...((((...((((.(((((--(--((((((((........)))).......))))))))))))))..)))) ( -19.11, z-score = -3.41, R) >droEre2.scaffold_4845 11521073 68 - 22589142 ACUGAAUUAAAGCUCAUAAG--C--AUGUUGGCCUAUAAACGCCAUUUUACAGCGUGUUUAUGGCUCUAUUU ...((((...(((.((((((--(--((((((((........)))).......))))))))))))))..)))) ( -19.41, z-score = -2.98, R) >consensus ACUGAAUUAAAGCCCAUAAGCUC__AUGUUGGCCUAUAAACGCCAUUUUACAGCGUGUUUAUGGCUCUAUUU ..........((((.((((((.(...(((((((........))))....)))..).))))))))))...... (-13.12 = -13.36 + 0.24)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:41:00 2011