
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 17,146,532 – 17,146,684 |
| Length | 152 |
| Max. P | 0.753872 |

| Location | 17,146,532 – 17,146,647 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 88.71 |
| Shannon entropy | 0.19179 |
| G+C content | 0.41823 |
| Mean single sequence MFE | -24.03 |
| Consensus MFE | -15.84 |
| Energy contribution | -15.96 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.668366 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 17146532 115 - 21146708 AACCUCUUGGCUGACACAGAAUUAUUCUUCUACUCCAGUGAUUCCAUUCGAAACUAUUUGACUGGAAUCAAUGGCGCCUCCUUUUUCUUU---UAGCAACUAUCAGAUUUUCCAACUC ........(((......((((......))))...(((.((((((((.(((((....))))).)))))))).))).))).......(((..---(((...)))..)))........... ( -23.90, z-score = -2.72, R) >droEre2.scaffold_4845 11290352 113 - 22589142 AAUCUCUGGACUGACAC--CAUUUUCCUUCUACUCCAGUGGUUCCCUUCAAAGCUGUUUGACUGGAAUCACUGGCGGCUCCUUCUUCCUC---GAGCAUCUAUCUGAUUUUCAAACUC ((((..((((.......--.............(.(((((((((((..(((((....)))))..))))))))))).)((((..........---)))).))))...))))......... ( -29.10, z-score = -2.88, R) >droYak2.chr2R 9096632 116 + 21139217 AACCUCUUGGCUGACAC--AAUUUUUCUUCUACUCCAGUGGUUGCCUUCAAAACUGUUUGACUGGAAUCAAUGGCGCCUCCUCCUUCCUCUUGGAGCAUCUAUCUGAUUUUCAAACUC ........(((...(((--..................)))...))).........((((((...((((((((((...((((...........))))...)))).)))))))))))).. ( -22.17, z-score = -0.95, R) >droSec1.super_9 476649 115 - 3197100 AACCUCUUGCCUGACACAGAAUUUUUCUUCUACUCCAGUGAUUCCAUUCGAAACUAUUUGACUGGAAUCAAUGGCGCCUCCUUCUUCUUU---UAGCAUCUAUCUGAUUUUCCAACUC ................((((..............(((.((((((((.(((((....))))).)))))))).))).((.............---..)).....))))............ ( -21.16, z-score = -2.36, R) >droSim1.chr2R 15797424 115 - 19596830 AACCUCUUGGCUGACACAGAAUUUUUCUUCUACUCCAGUGAUUCCAUUCGAAACUAUUUGACUGGAAUCAAUGGCGCCUCCUUCUUCUUU---UAGCAUCUAUCUGAUUUUCCAACUC .........(((((...((((......)))).(.(((.((((((((.(((((....))))).)))))))).))).).............)---))))..................... ( -23.80, z-score = -2.76, R) >consensus AACCUCUUGGCUGACACAGAAUUUUUCUUCUACUCCAGUGAUUCCAUUCGAAACUAUUUGACUGGAAUCAAUGGCGCCUCCUUCUUCUUU___UAGCAUCUAUCUGAUUUUCCAACUC ................................(.(((.((((((((.(((((....))))).)))))))).))).).......................................... (-15.84 = -15.96 + 0.12)



| Location | 17,146,567 – 17,146,684 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 88.44 |
| Shannon entropy | 0.20498 |
| G+C content | 0.43050 |
| Mean single sequence MFE | -26.14 |
| Consensus MFE | -17.66 |
| Energy contribution | -17.74 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.753872 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 17146567 117 - 21146708 AUCUCUGCUAACUUCGCAGAUUAUGCUCAAAUUAACCAACCUCUUGGCUGACACAGAAUUAUUCUUCUACUCCAGUGAUUCCAUUCGAAACUAUUUGACUGGAAUCAAUGGCGCCUC ...(((((.......))))).........................(((......((((......))))...(((.((((((((.(((((....))))).)))))))).))).))).. ( -28.80, z-score = -3.88, R) >droEre2.scaffold_4845 11290387 115 - 22589142 AUCUCUCCUAAUUUCGCAGAUCGUGUCAGAAUCGACCAAUCUCUGGACUGACAC--CAUUUUCCUUCUACUCCAGUGGUUCCCUUCAAAGCUGUUUGACUGGAAUCACUGGCGGCUC ...............((((((.(((((((......(((.....))).)))))))--.))))........(.(((((((((((..(((((....)))))..))))))))))).))).. ( -34.70, z-score = -3.27, R) >droYak2.chr2R 9096670 115 + 21139217 AUCUCACCUAAUUUCGCAGAUCCUGGCAAAAUUGACCAACCUCUUGGCUGACAC--AAUUUUUCUUCUACUCCAGUGGUUGCCUUCAAAACUGUUUGACUGGAAUCAAUGGCGCCUC ........................((((((((((..((.((....)).))...)--)))))).........(((.(((((.((.(((((....)))))..))))))).))).))).. ( -21.30, z-score = -0.03, R) >droSec1.super_9 476684 117 - 3197100 AUCUCUCCUAAUUUCGCAGAUCAUGACCAAAUUAACCAACCUCUUGCCUGACACAGAAUUUUUCUUCUACUCCAGUGAUUCCAUUCGAAACUAUUUGACUGGAAUCAAUGGCGCCUC ........((((((.(((.....)).).))))))...........(((......((((......)))).......((((((((.(((((....))))).))))))))..)))..... ( -21.20, z-score = -1.85, R) >droSim1.chr2R 15797459 117 - 19596830 AUCUCUCCUAAUUUCGCAGAUCAUGACCAAAUUAACCAACCUCUUGGCUGACACAGAAUUUUUCUUCUACUCCAGUGAUUCCAUUCGAAACUAUUUGACUGGAAUCAAUGGCGCCUC ........((((((.(((.....)).).))))))...........(((......((((......))))...(((.((((((((.(((((....))))).)))))))).))).))).. ( -24.70, z-score = -2.44, R) >consensus AUCUCUCCUAAUUUCGCAGAUCAUGACCAAAUUAACCAACCUCUUGGCUGACACAGAAUUUUUCUUCUACUCCAGUGAUUCCAUUCGAAACUAUUUGACUGGAAUCAAUGGCGCCUC ..............(((...................((.((....)).))......................((.((((((((.(((((....))))).)))))))).))))).... (-17.66 = -17.74 + 0.08)

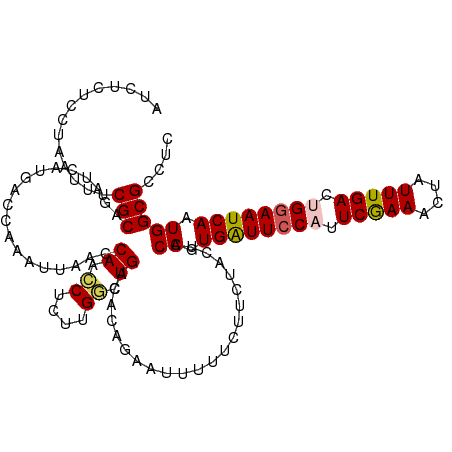

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:40:25 2011