
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 16,555,258 – 16,555,368 |
| Length | 110 |
| Max. P | 0.623520 |

| Location | 16,555,258 – 16,555,368 |
|---|---|
| Length | 110 |
| Sequences | 11 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 64.72 |
| Shannon entropy | 0.72524 |
| G+C content | 0.41188 |
| Mean single sequence MFE | -23.27 |
| Consensus MFE | -8.64 |
| Energy contribution | -9.26 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.08 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.623520 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 16555258 110 - 21146708 -UGUACAA---UUAAGAACU---CGCUGCGUUUAGGCAUAGUUUUCCCGGCGA-ACCGAUGCA-CAUCGCCGCACUCAAAUGGGAGAUUGGGAAAAUAUGAUCCAAAACGAAACGCCUC -.......---.........---.((..(((((.((((((.((((((((((((-.........-..)))))...(((......)))...)))))))))))..)).)))))....))... ( -28.30, z-score = -0.96, R) >droSec1.super_1 14109229 109 - 14215200 -UGUACAA---UUAAGAACU---CGCUGCA-UUAAGCAUAGUUUUCCAGGCGA-ACCGAUGCA-CAUCGCCGAACUCAAAUGAGAGAUUGGAAAAAUAUGAUUCAAAACGGAACGCCUC -.......---...((((((---.(((...-...)))..))))))..(((((.-.(((((...-.))).((((.(((......))).))))..................))..))))). ( -23.70, z-score = -1.18, R) >droSim1.chr2R 15216513 109 - 19596830 -UGUACAA---UUAAGAACU---CGCUGCA-UUAAGCAUAGUUUUCCAGGCGA-ACCGAUGCA-CAUCGCCGAACUCAAAUGAGAGAUUGGAAAAAUAUGAUUCAAAACGGAACGCCUC -.......---...((((((---.(((...-...)))..))))))..(((((.-.(((((...-.))).((((.(((......))).))))..................))..))))). ( -23.70, z-score = -1.18, R) >droAna3.scaffold_13266 15856918 112 + 19884421 UUGUACAGAGUUCAAUAAC----CGCAGCGUUUAGGCAUAGUUUUCCAUGCGG-ACCGAUGCACCGAUGUCGUACCAGGAUGGGCGAUUGGAAAAAUAUGAUCCAUA--AAAACGCCUC ...................----....((((((.((((((.(((((((((((.-.....)))).....(((((.((.....)))))))))))))).))))..))...--.))))))... ( -24.70, z-score = 0.52, R) >droEre2.scaffold_4845 10703346 111 - 22589142 -UGUACAA---UGAGGAACUGGCUGCUGCGUUUAGGCAUAGUUUUCCAUGCGA-AGCGAUGCA-CAAAGUCGCACCAGAAUGGGCGAUUGGAAAAAUAUGAUCCAAA--GAAACGCCUC -.......---.((((.....((....))((((.((((((.((((((.((((.-.....))))-...((((((.((.....)))))))))))))).))))..))...--.)))).)))) ( -29.20, z-score = -0.58, R) >droYak2.chr2R 8498759 111 + 21139217 --AUACAA--UUAAAGAACUCGCUGCUGCGUUUAGGCAUAGUUUUCCAUGCGA-ACCGAUGCA-CAAAGUCGCACCAAAAAGAGCGAUUGGAAAAAUAUGAUCCAAA--GAAACGCCUU --......--.................((((((.((((((.((((((.((((.-.....))))-...((((((.(......).)))))))))))).))))..))...--.))))))... ( -24.00, z-score = -1.04, R) >droPer1.super_4 4684809 100 + 7162766 -------C---GUACAAAU----CGUCGCGUUUAGACAUAGUUUUCCAAAAAAUA-CGAUGUACAACUGUACUGUCGGA--GGGCCAUUGGAAAAAUAUGAUCCCAA--UAAACGCCUC -------.---........----....(((((((((((((.((((((((......-.(((((((....)))).)))((.--...)).)))))))).)))).))....--)))))))... ( -25.40, z-score = -2.42, R) >dp4.chr3 11186067 100 + 19779522 -------C---GUACAAAU----CGUCGCGUUUAGACAUAGUUUUCCAAAAAAUA-CGAUGUACAACUGUACUGUCGGA--GGGCCAUUGGAAAAAUAUGAUCCCAA--UAAACGCCUC -------.---........----....(((((((((((((.((((((((......-.(((((((....)))).)))((.--...)).)))))))).)))).))....--)))))))... ( -25.40, z-score = -2.42, R) >droWil1.scaffold_180700 4548015 108 - 6630534 UAACAAUA---UGAUCAAU----CGUUGCGUUUAGACAUAGUUUUCCAUUUGUUAGCUCUGCAACUAAAAGAAAAAAAA--GAACGAUUGGAAAAAUAUGAGUCAGA--UAAACGCUUC ........---........----....((((((((((.((.(((((((.(((((...(((.........))).......--.))))).)))))))...)).)))...--)))))))... ( -20.40, z-score = -1.48, R) >droVir3.scaffold_12823 260759 93 + 2474545 -UAUGCUA---UCAAAUAU----CGUUGCGUUCAGGCAUAGUUUUCCA--------------AAAUAUAUGAAUUACGG--ACACGAUUGGAAAAAUAUGUACCCCA--GAAACGCCUC -.......---........----....((((((.((((((.(((((((--------------((((......))).((.--...)).)))))))).))))....)).--).)))))... ( -15.50, z-score = -0.19, R) >droGri2.scaffold_15245 15865474 90 - 18325388 -UAUGCCA---UAAAAUAU----CGUUGCGUUCAGGUAUAGUUUUCC----------------AGCAAAUCAUGUGCAA--CAACGCUUGGAAAAAUAU-UAACCAA--AAAACGCUUC -.......---........----....(((((..((((((.((((((----------------((((.....)))((..--....)).))))))).)))-..)))..--..)))))... ( -15.70, z-score = -1.01, R) >consensus _UGUACAA___UUAAGAAC____CGCUGCGUUUAGGCAUAGUUUUCCAUGCGA_ACCGAUGCA_CAAAGUCGAACCCAAA_GAGCGAUUGGAAAAAUAUGAUCCAAA__GAAACGCCUC ...........................((((((...((((.(((((((........................................))))))).))))..........))))))... ( -8.64 = -9.26 + 0.62)

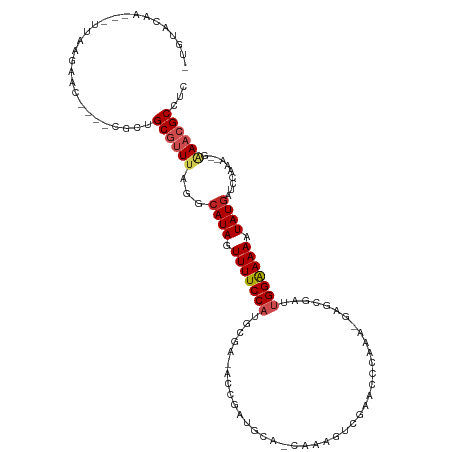
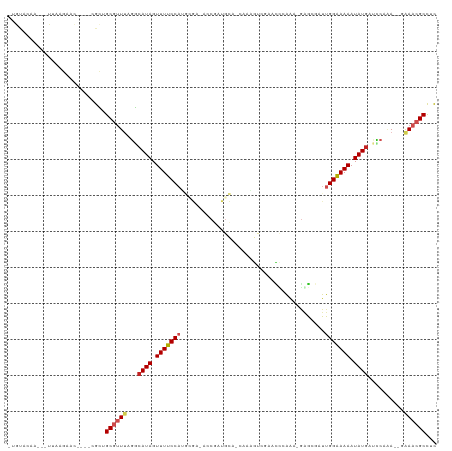
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:38:53 2011