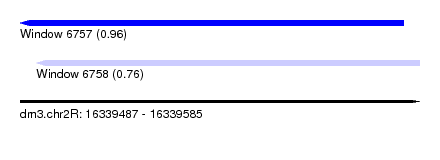

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 16,339,487 – 16,339,585 |
| Length | 98 |
| Max. P | 0.960987 |

| Location | 16,339,487 – 16,339,581 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 90.35 |
| Shannon entropy | 0.18445 |
| G+C content | 0.53916 |
| Mean single sequence MFE | -35.42 |
| Consensus MFE | -27.22 |
| Energy contribution | -27.37 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.69 |
| SVM RNA-class probability | 0.960987 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
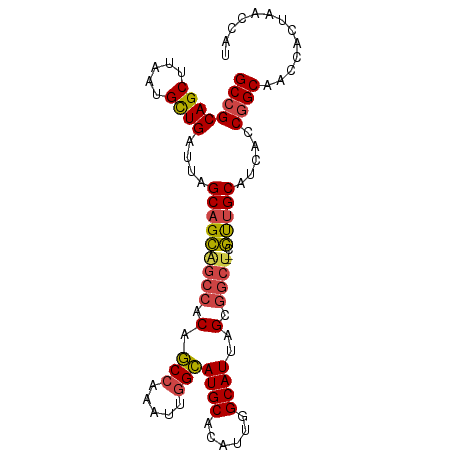
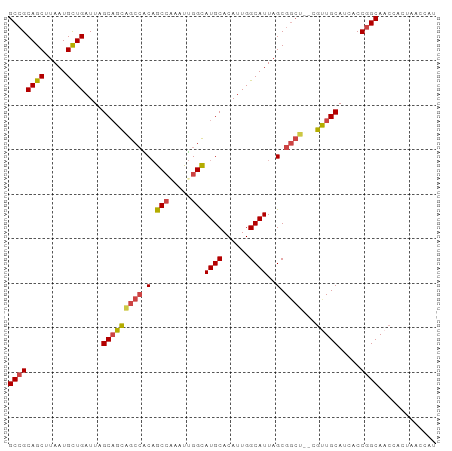
>dm3.chr2R 16339487 94 - 21146708 GCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCCAAAUUGGCAUGCACAUUGGCAUUAGCGGCU--CGUUGCAUCACCGGCAACCACUAACCAU ((((((((.....))))....(((((((((.(.(((.....)))((((......))))..).))))--.))))).....))))............. ( -34.80, z-score = -2.67, R) >droAna3.scaffold_13266 16933633 96 + 19884421 GCGGCAGCUUAAUGUUGAUUAGCUGUCGCAGCCACAGGAUCGGUAUGCACAUUGGCAUUAGCGAAUACCACUGCAACACCGGCAACCGCUAACCAU (((((((((.(((....))))))))))))(((.....(.(((((.((((...(((.(((....))).))).))))..))))))....)))...... ( -29.20, z-score = -0.97, R) >droEre2.scaffold_4845 10485379 93 - 22589142 GCCGCAGCUUAAUGCUGAUUAGCAGCGGCCGCAGCC-AAUUGGCAUGCACAUUGGCAUUAGCGGCU--CGUUGCACCACCGGCAACCACUAACCAU ((((((((.....))))....(((((((((((.(((-....)))((((......))))..))))).--)))))).....))))............. ( -41.70, z-score = -3.79, R) >droYak2.chr2R 8280287 93 + 21139217 GCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCC-AAUUGGCAUGCACAUUGGCAUUAGCGGCU--CGCUGCACCACCGGCAACCACUAACCAU ((((((((.....))))....(((((((((.(.(((-....)))((((......))))..).))))--.))))).....))))............. ( -37.20, z-score = -3.22, R) >droSec1.super_1 13896067 94 - 14215200 GCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCCAAAUUGGCAUGCACAUUGGCAUUAGCGGCU--AGUUGCAUCACCGGCAACCACUAACCAU ((((((((.....))))....(((((((((.(.(((.....)))((((......))))..).))))--.))))).....))))............. ( -34.80, z-score = -2.61, R) >droSim1.chr2R 14996374 94 - 19596830 GCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCCAAAUUGGCAUGCACAUUGGCAUUAGCGGCU--CGUUGCAUCACCGGCAACCACUAACCAU ((((((((.....))))....(((((((((.(.(((.....)))((((......))))..).))))--.))))).....))))............. ( -34.80, z-score = -2.67, R) >consensus GCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCCAAAUUGGCAUGCACAUUGGCAUUAGCGGCU__CGUUGCAUCACCGGCAACCACUAACCAU ((((((((.....))))....(((((((((.(.(((.....)))((((......))))..).))))...))))).....))))............. (-27.22 = -27.37 + 0.14)

| Location | 16,339,491 – 16,339,585 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 90.35 |
| Shannon entropy | 0.18445 |
| G+C content | 0.52852 |
| Mean single sequence MFE | -36.17 |
| Consensus MFE | -28.12 |
| Energy contribution | -28.27 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.756473 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
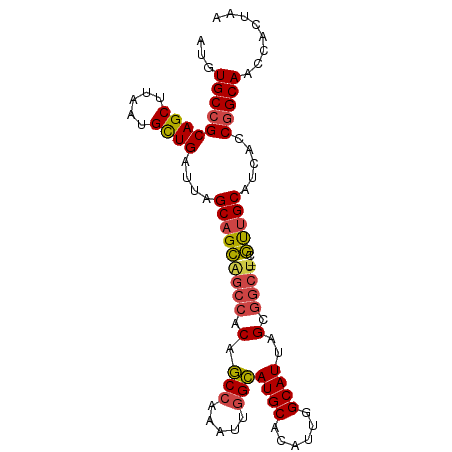
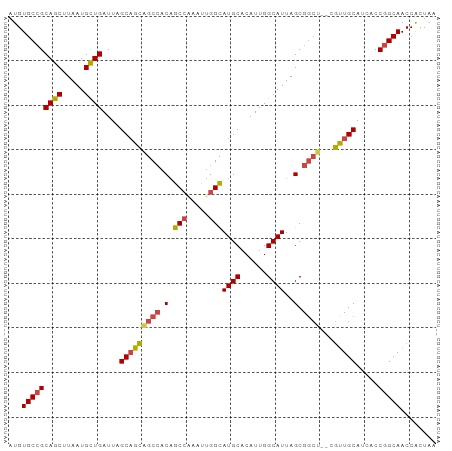
>dm3.chr2R 16339491 94 - 21146708 AUGUGCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCCAAAUUGGCAUGCACAUUGGCAUUAGCGGCU--CGUUGCAUCACCGGCAACCACUAA ...(((((((((.....))))....(((((((((.(.(((.....)))((((......))))..).))))--.))))).....)))))........ ( -35.70, z-score = -1.99, R) >droAna3.scaffold_13266 16933637 96 + 19884421 AUGUGCGGCAGCUUAAUGUUGAUUAGCUGUCGCAGCCACAGGAUCGGUAUGCACAUUGGCAUUAGCGAAUACCACUGCAACACCGGCAACCGCUAA ....(((((((((.(((....))))))))))))(((.....(.(((((.((((...(((.(((....))).))).))))..))))))....))).. ( -29.20, z-score = -0.26, R) >droEre2.scaffold_4845 10485383 93 - 22589142 AUGUGCCGCAGCUUAAUGCUGAUUAGCAGCGGCCGCAGCC-AAUUGGCAUGCACAUUGGCAUUAGCGGCU--CGUUGCACCACCGGCAACCACUAA ...(((((((((.....))))....(((((((((((.(((-....)))((((......))))..))))).--)))))).....)))))........ ( -42.60, z-score = -3.20, R) >droYak2.chr2R 8280291 93 + 21139217 AUGUGCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCC-AAUUGGCAUGCACAUUGGCAUUAGCGGCU--CGCUGCACCACCGGCAACCACUAA ...(((((((((.....))))....(((((((((.(.(((-....)))((((......))))..).))))--.))))).....)))))........ ( -38.10, z-score = -2.47, R) >droSec1.super_1 13896071 94 - 14215200 AUGUGCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCCAAAUUGGCAUGCACAUUGGCAUUAGCGGCU--AGUUGCAUCACCGGCAACCACUAA ...(((((((((.....))))....(((((((((.(.(((.....)))((((......))))..).))))--.))))).....)))))........ ( -35.70, z-score = -1.95, R) >droSim1.chr2R 14996378 94 - 19596830 AUGUGCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCCAAAUUGGCAUGCACAUUGGCAUUAGCGGCU--CGUUGCAUCACCGGCAACCACUAA ...(((((((((.....))))....(((((((((.(.(((.....)))((((......))))..).))))--.))))).....)))))........ ( -35.70, z-score = -1.99, R) >consensus AUGUGCCGCAGCUUAAUGCUGAUUAGCAGCAGCCACAGCCAAAUUGGCAUGCACAUUGGCAUUAGCGGCU__CGUUGCAUCACCGGCAACCACUAA ...(((((((((.....))))....(((((((((.(.(((.....)))((((......))))..).))))...))))).....)))))........ (-28.12 = -28.27 + 0.14)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:38:27 2011