
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 389,003 – 389,115 |
| Length | 112 |
| Max. P | 0.737449 |

| Location | 389,003 – 389,095 |
|---|---|
| Length | 92 |
| Sequences | 11 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 83.21 |
| Shannon entropy | 0.35067 |
| G+C content | 0.36471 |
| Mean single sequence MFE | -20.71 |
| Consensus MFE | -14.20 |
| Energy contribution | -14.28 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.737449 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 389003 92 - 23011544 --GUUUAAU--CAAAGUGGCGCUAAUUAGUUUGA---GUCAUUUGGAUAAUUGCAAAUUAAACGCAAACUUAAUUGCGCCAACACACGAAACACACACA---------- --((((...--.....((((((.((((((((((.---((.(((((.(....).)))))...)).))))).))))))))))).......)))).......---------- ( -24.26, z-score = -3.61, R) >droSim1.chr2L 395464 92 - 22036055 --GUUUAAU--CAAAGUGGCGCUAAUUAGUUUGA---GUCGUUUGGAUAAUUGCAAAUUAAACGCAAACUUAAUUGCGCCAACACACGAAACACACAUA---------- --((((...--.....((((((.((((((((((.---((.(((((.(....).)))))...)).))))).))))))))))).......)))).......---------- ( -24.56, z-score = -3.20, R) >droSec1.super_14 372510 92 - 2068291 --GUUUAAU--CAAAGUGGCGCUAAUUAGUUUGA---GUCGUUUGGAUAAUUGCAAAUUAAACGCAAACUUAAUUGCGCCAACACACGAAACACACAUA---------- --((((...--.....((((((.((((((((((.---((.(((((.(....).)))))...)).))))).))))))))))).......)))).......---------- ( -24.56, z-score = -3.20, R) >droYak2.chr2L 372577 102 - 22324452 --GUUUAAU--CAAAGUGGCGCUAAUUAGUUCGA---GUCGUUUGGAUAAUUGCAAAUUAAACGCAAACUUAAUUGCGCCAACACACGAAUCGCACACACACUCGCACA --.......--....(((.((.......((.(((---.((((((((.((((.....))))..(((((......)))))))))...)))).))).)).......))))). ( -18.54, z-score = 0.20, R) >droEre2.scaffold_4929 439648 88 - 26641161 --GUUUAAU--CAAAGUGGCGCUAAUUAGUUCGA---GUCGUUUGGAUAAUUGCAAAUUAAACGCAAACUUAAUUGCGCCAACACACGAAACACA-------------- --((((...--.....((((((.((((((((.(.---((.(((((.(....).)))))...)).).))).))))))))))).......))))...-------------- ( -20.46, z-score = -1.84, R) >droAna3.scaffold_12943 170879 92 + 5039921 --GAUUAAU--CAAAGUGGCGCUAAUUAGUUUGA---GUCGUUUGGAUAAUUGCAAAUUAAACGCAAACUUAAUUGCGCCGACACACGAAACACACACA---------- --.......--....(((((((.((((((((((.---((.(((((.(....).)))))...)).))))).)))))))))).))................---------- ( -21.80, z-score = -2.17, R) >dp4.chr4_group3 10443161 98 + 11692001 --GCUUAAU--CAAAGUGGCGCUAAUUAGUUUGA---GUCGUUUGGAUAAUUGCAAAUUAAACGCAAACUUAAUUGCGCCGACACACGAAACACACAGAUGCACA---- --.......--....(((((((.((((((((((.---((.(((((.(....).)))))...)).))))).)))))))))).))......................---- ( -21.80, z-score = -0.94, R) >droWil1.scaffold_180703 2560078 92 + 3946847 --GUUUAAU--CAAAGUGGCGCUAAUUAGUUUGA---GUCGUUUGGAUAAUUGCAAAUUAAACGCAAAGUUAAUUGCACCGACACACGAAACACACAUA---------- --((((...--.....(((.((.((((((((((.---((.(((((.(....).)))))...)).)))).)))))))).))).......)))).......---------- ( -16.16, z-score = -0.04, R) >droVir3.scaffold_12963 18781127 92 - 20206255 --GUUUAAU--CAAAGUGGCGCUAAUUAGUUUGA---GUCGUUUGGAUAAUUGCAAAUUAAACGCAAACUUAAUUGCGCCGACACACGAAACACACACA---------- --((((...--.....((((((.((((((((((.---((.(((((.(....).)))))...)).))))).))))))))))).......)))).......---------- ( -23.86, z-score = -2.57, R) >droGri2.scaffold_15252 6702819 92 - 17193109 --GUUAAAU--CAAAUUGGCGCUAAUUAGUUUGA---GUCGUUUGGAUAAUUGCAAAUUAAACGCAAAGUUAAUUGCGCCGACACACGAAACACACAUA---------- --.......--....(((((((.((((((((((.---((.(((((.(....).)))))...)).)))).))))))))))))).................---------- ( -20.10, z-score = -1.30, R) >triCas2.ChLG8 632744 99 - 15773733 UUGUCAAAUUGCAAGGCUACUAUGAUUUUUUGGAAAUGAUGAUUAAAAAAUUAUUAUAAAACAGUUUUUUCAAAUGGAAUGACCCUGGUUUUUCACUGA---------- (((..((((((........(((........)))..(((((((((....)))))))))....))))))...)))...(((.(((....))).))).....---------- ( -11.70, z-score = 1.34, R) >consensus __GUUUAAU__CAAAGUGGCGCUAAUUAGUUUGA___GUCGUUUGGAUAAUUGCAAAUUAAACGCAAACUUAAUUGCGCCGACACACGAAACACACACA__________ ................((((((.((((((((((.......(((((.(....).)))))......)))).))))))))))))............................ (-14.20 = -14.28 + 0.08)


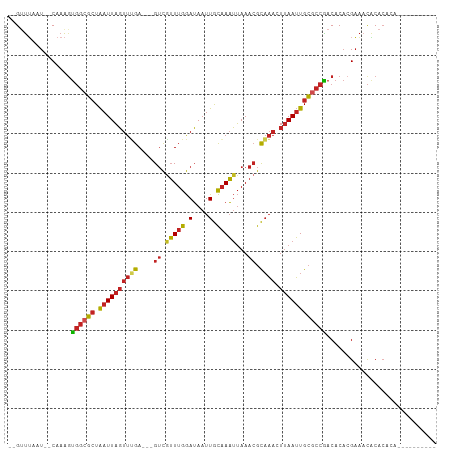
| Location | 389,022 – 389,115 |
|---|---|
| Length | 93 |
| Sequences | 12 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 89.24 |
| Shannon entropy | 0.23204 |
| G+C content | 0.34896 |
| Mean single sequence MFE | -18.69 |
| Consensus MFE | -14.46 |
| Energy contribution | -14.25 |
| Covariance contribution | -0.21 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.670707 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr2L 389022 93 + 23011544 GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAAUGACUCAAACUAAUUAGCGCCACUUUGAUUAAACUUAGCAGCACAAAAAUUGAA ((((((((((.(((((.(((..(((((........)))))))).)))))))))).)))))................................. ( -20.90, z-score = -2.13, R) >droSim1.chr2L 395483 93 + 22036055 GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCACUUUGAUUAAACUUAGCAGCACAAAAAUUGAA ((((((((((.(((((.(((...((((........)))).))).)))))))))).)))))................................. ( -19.50, z-score = -1.80, R) >droSec1.super_14 372529 93 + 2068291 GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCACUUUGAUUAAACUUAGCAGCACAAAAAUUGAA ((((((((((.(((((.(((...((((........)))).))).)))))))))).)))))................................. ( -19.50, z-score = -1.80, R) >droYak2.chr2L 372606 92 + 22324452 GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCGAACUAAUUAGCGCCACUUUGAUUAAACUUAGCAGCA-CAAAAUUGAA ((((((((((.(((((.(((...((((........)))).))).)))))))))).)))))......................-.......... ( -19.20, z-score = -1.47, R) >droEre2.scaffold_4929 439663 92 + 26641161 GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCGAACUAAUUAGCGCCACUUUGAUUAAACUUAGCAGCA-CAAAAUUGAA ((((((((((.(((((.(((...((((........)))).))).)))))))))).)))))......................-.......... ( -19.20, z-score = -1.47, R) >droAna3.scaffold_12943 170898 92 - 5039921 GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCACUUUGAUUAAUCCUGGCAACAACAAA-UGGAA ((((((((((.(((((.(((...((((........)))).))).)))))))))).)))))...........(((((....)).....-.))). ( -22.30, z-score = -2.75, R) >droPer1.super_1 7562929 93 - 10282868 GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCACUUUGAUUAAGCCUGCCAACAACAAAAUGGAA ((((((((((.(((((.(((...((((........)))).))).)))))))))).)))))................(((.........))).. ( -20.10, z-score = -1.73, R) >dp4.chr4_group3 10443186 93 - 11692001 GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCACUUUGAUUAAGCCUGCCAACAACAAAAUGGAA ((((((((((.(((((.(((...((((........)))).))).)))))))))).)))))................(((.........))).. ( -20.10, z-score = -1.73, R) >droWil1.scaffold_180703 2560097 83 - 3946847 GGUGCAAUUAACUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCACUUUGAUUAAACUUGGCAAGAA---------- ((((((((((..((((.(((...((((........)))).))).)))).))))).))))).......................---------- ( -13.40, z-score = -0.52, R) >droVir3.scaffold_12963 18781146 87 + 20206255 GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCACUUUGAUUAAACUUUGCAAAAUUGAA------ ((((((((((.(((((.(((...((((........)))).))).)))))))))).)))))...........................------ ( -19.50, z-score = -2.32, R) >droMoj3.scaffold_6500 7969044 87 - 32352404 GGCGCAAUUAACUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCACUUUGAUUAAACUUUGCAAAAUUGAA------ ((((((((((..((((.(((...((((........)))).))).)))).))))).)))))...........................------ ( -15.30, z-score = -1.36, R) >droGri2.scaffold_15252 6702838 87 + 17193109 GGCGCAAUUAACUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCAAUUUGAUUUAACUUUGCAAAAUUUAA------ ((((((((((..((((.(((...((((........)))).))).)))).))))).)))))...........................------ ( -15.30, z-score = -1.43, R) >consensus GGCGCAAUUAAGUUUGCGUUUAAUUUGCAAUUAUCCAAACGACUCAAACUAAUUAGCGCCACUUUGAUUAAACUUAGCAACAACAAAAUUGAA ((((((((((...(((.(((...((((........)))).))).)))..))))).)))))................................. (-14.46 = -14.25 + -0.21)


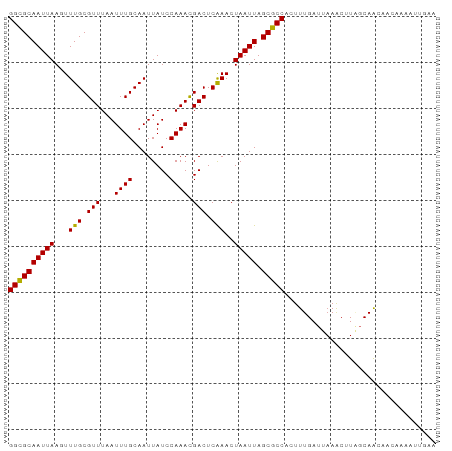
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:05:41 2011