
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 16,123,918 – 16,124,054 |
| Length | 136 |
| Max. P | 0.976847 |

| Location | 16,123,918 – 16,124,015 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 92.85 |
| Shannon entropy | 0.14023 |
| G+C content | 0.61725 |
| Mean single sequence MFE | -42.10 |
| Consensus MFE | -39.79 |
| Energy contribution | -39.35 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.832898 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
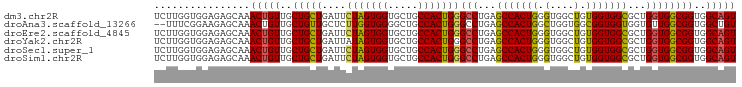
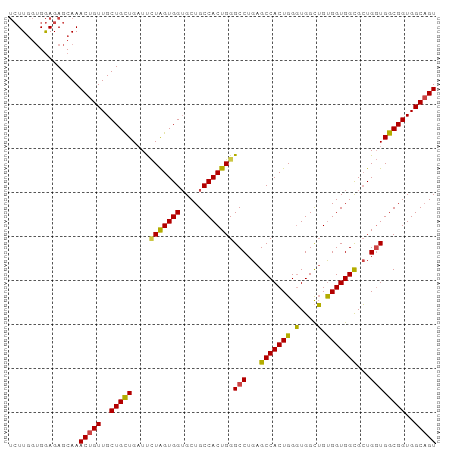
>dm3.chr2R 16123918 97 + 21146708 UCUUGGUGGAGAGCAAACUGUUGCUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGU ((((....))))....(((((..(((((....(((((((.....)))))))(((...(((((((.(....).)))))))...))))))))..))))) ( -43.10, z-score = -1.50, R) >droAna3.scaffold_13266 10665352 95 - 19884421 --UUUCGGAAGAGCAAACUGUUGCUGUUGCUCUUGGUGGGGCUGCCACUGGGCCUGAGCCACUGGCUGGUGGCGGUGGUGGUGUUGGCGGUGGCUGU --.(((....)))...((.((..((((((((((....))))).((((((..(((...((((((....)))))))))))))))...)))))..)).)) ( -38.20, z-score = -0.63, R) >droEre2.scaffold_4845 10274274 97 + 22589142 UCUUGGUGGAGAGCAAACUGUUGCUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGU ((((....))))....(((((..(((((....(((((((.....)))))))(((...(((((((.(....).)))))))...))))))))..))))) ( -43.10, z-score = -1.50, R) >droYak2.chr2R 8059458 97 - 21139217 UCUUGGUGGAGAGCAAACUGUUGCUGCUGAUUAUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGU ..(..((((..((((.(((((..(....)...)))))..)))).))))..)(((...(((((..(((.((....)).)).)..)))))...)))... ( -42.00, z-score = -1.51, R) >droSec1.super_1 13690168 97 + 14215200 UCUUGGUGGAGAGCAAACUGUUGCUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGU ((((....))))....(((((..(((((....(((((((.....)))))))(((...(((((((.(....).)))))))...))))))))..))))) ( -43.10, z-score = -1.50, R) >droSim1.chr2R 14782288 97 + 19596830 UCUUGGUGGAGAGCAAACUGUUGCUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGU ((((....))))....(((((..(((((....(((((((.....)))))))(((...(((((((.(....).)))))))...))))))))..))))) ( -43.10, z-score = -1.50, R) >consensus UCUUGGUGGAGAGCAAACUGUUGCUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGU ................(((((..(((((....(((((((.....)))))))(((...(((((((.(....).)))))))...))))))))..))))) (-39.79 = -39.35 + -0.44)

| Location | 16,123,918 – 16,124,015 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 92.85 |
| Shannon entropy | 0.14023 |
| G+C content | 0.61725 |
| Mean single sequence MFE | -27.57 |
| Consensus MFE | -22.90 |
| Energy contribution | -23.40 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.515634 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
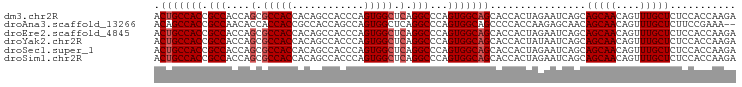
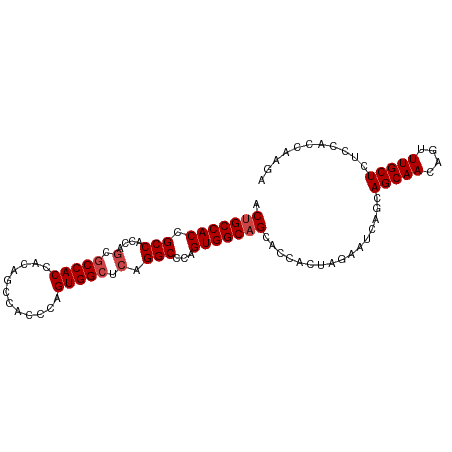
>dm3.chr2R 16123918 97 - 21146708 ACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAGCAACAGUUUGCUCUCCACCAAGA .(((((((.((.....))(((.....((((((...))))))..)))...))))))).............((.(((((....)))))))......... ( -27.80, z-score = -1.39, R) >droAna3.scaffold_13266 10665352 95 + 19884421 ACAGCCACCGCCAACACCACCACCGCCACCAGCCAGUGGCUCAGGCCCAGUGGCAGCCCCACCAAGAGCAACAGCAACAGUUUGCUCUUCCGAAA-- ...(((((.(((....((((....((.....))..))))....)))...))))).........((((((((..((....))))))))))......-- ( -26.40, z-score = -2.10, R) >droEre2.scaffold_4845 10274274 97 - 22589142 ACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAGCAACAGUUUGCUCUCCACCAAGA .(((((((.((.....))(((.....((((((...))))))..)))...))))))).............((.(((((....)))))))......... ( -27.80, z-score = -1.39, R) >droYak2.chr2R 8059458 97 + 21139217 ACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAUAAUCAGCAGCAACAGUUUGCUCUCCACCAAGA .(((((((.((.....))(((.....((((((...))))))..)))...))))))).............((.(((((....)))))))......... ( -27.80, z-score = -1.87, R) >droSec1.super_1 13690168 97 - 14215200 ACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAGCAACAGUUUGCUCUCCACCAAGA .(((((((.((.....))(((.....((((((...))))))..)))...))))))).............((.(((((....)))))))......... ( -27.80, z-score = -1.39, R) >droSim1.chr2R 14782288 97 - 19596830 ACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAGCAACAGUUUGCUCUCCACCAAGA .(((((((.((.....))(((.....((((((...))))))..)))...))))))).............((.(((((....)))))))......... ( -27.80, z-score = -1.39, R) >consensus ACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAGCAACAGUUUGCUCUCCACCAAGA .(((((((.(((....(.(((((............))))).).)))...)))))))................(((((....)))))........... (-22.90 = -23.40 + 0.50)
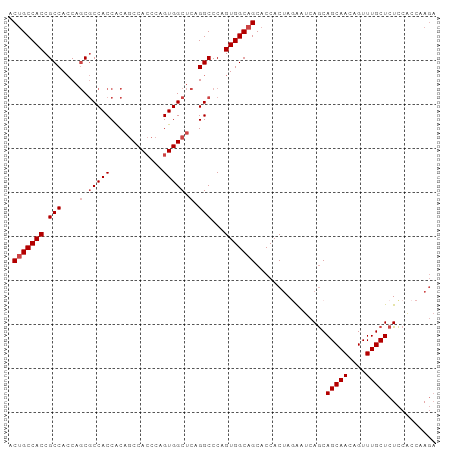
| Location | 16,123,941 – 16,124,054 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 99.29 |
| Shannon entropy | 0.01278 |
| G+C content | 0.60708 |
| Mean single sequence MFE | -48.28 |
| Consensus MFE | -46.44 |
| Energy contribution | -46.44 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.96 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.795965 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 16123941 113 + 21146708 CUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGUUUUUGUUUGGCGGGCGGCGGUGCCAAAUGAUGAUCAUGA ....((((((...(((..(((((.((..(((...(((((..(((.((....)).)).)..)))))...))).(((.......))).)).)))))..)))...))..))))... ( -46.80, z-score = -0.95, R) >droEre2.scaffold_4845 10274297 113 + 22589142 CUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGUUUUUGUUUGGCGGGCGGCGGUGCCAAAUGAUGAUCAUGA ....((((((...(((..(((((.((..(((...(((((..(((.((....)).)).)..)))))...))).(((.......))).)).)))))..)))...))..))))... ( -46.80, z-score = -0.95, R) >droYak2.chr2R 8059481 113 - 21139217 CUGCUGAUUAUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGUUUUUGUUUGGCGGGUGGCGGUGCCAAAUGAUGAUCAUGA ....(((((((..(((..((((((((..(((...(((((..(((.((....)).)).)..)))))...))).(((.......))).))))))))..)))....)))))))... ( -54.20, z-score = -3.94, R) >droSec1.super_1 13690191 113 + 14215200 CUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGUUUUUGUUUGGCGGGCGGCGGUGCCAAAUGAUGAUCAUGA ....((((((...(((..(((((.((..(((...(((((..(((.((....)).)).)..)))))...))).(((.......))).)).)))))..)))...))..))))... ( -46.80, z-score = -0.95, R) >droSim1.chr2R 14782311 113 + 19596830 CUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGUUUUUGUUUGGCGGGCGGCGGUGCCAAAUGAUGAUCAUGA ....((((((...(((..(((((.((..(((...(((((..(((.((....)).)).)..)))))...))).(((.......))).)).)))))..)))...))..))))... ( -46.80, z-score = -0.95, R) >consensus CUGCUGAUUCUAGUGGUGCUGCCACUGGGCCUGAGCCACUGGGUGGCUGUGGUGGCGCUGGUGGCGGUGGCAGUUUUUGUUUGGCGGGCGGCGGUGCCAAAUGAUGAUCAUGA ....(((((....(((..(((((.((..(((...(((((..(((.((....)).)).)..)))))...))).(((.......))).)).)))))..)))......)))))... (-46.44 = -46.44 + 0.00)

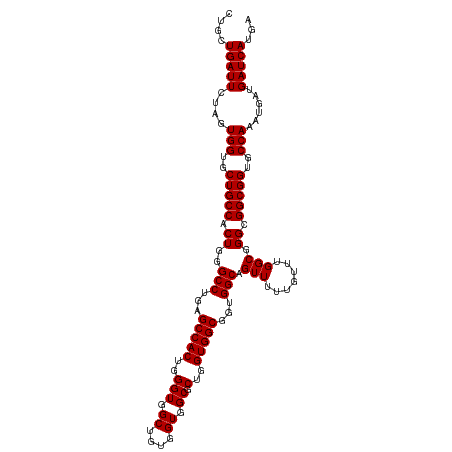
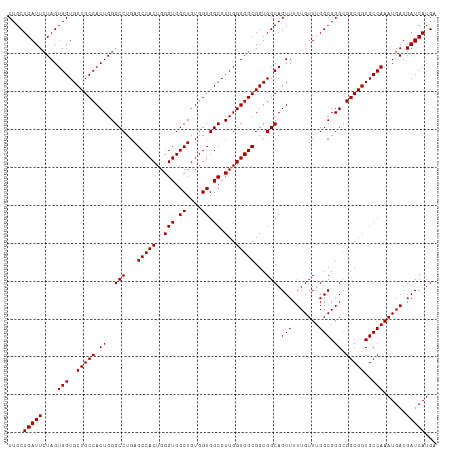
| Location | 16,123,941 – 16,124,054 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 99.29 |
| Shannon entropy | 0.01278 |
| G+C content | 0.60708 |
| Mean single sequence MFE | -31.98 |
| Consensus MFE | -31.50 |
| Energy contribution | -31.50 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.98 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.96 |
| SVM RNA-class probability | 0.976847 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 16123941 113 - 21146708 UCAUGAUCAUCAUUUGGCACCGCCGCCCGCCAAACAAAAACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAG ...(((((....((((((..........))))))......(((((((.((.....))(((.....((((((...))))))..)))...)))))))........).)))).... ( -32.10, z-score = -2.16, R) >droEre2.scaffold_4845 10274297 113 - 22589142 UCAUGAUCAUCAUUUGGCACCGCCGCCCGCCAAACAAAAACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAG ...(((((....((((((..........))))))......(((((((.((.....))(((.....((((((...))))))..)))...)))))))........).)))).... ( -32.10, z-score = -2.16, R) >droYak2.chr2R 8059481 113 + 21139217 UCAUGAUCAUCAUUUGGCACCGCCACCCGCCAAACAAAAACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAUAAUCAGCAG ...((((.....((((((..........))))))......(((((((.((.....))(((.....((((((...))))))..)))...)))))))..........)))).... ( -31.50, z-score = -2.79, R) >droSec1.super_1 13690191 113 - 14215200 UCAUGAUCAUCAUUUGGCACCGCCGCCCGCCAAACAAAAACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAG ...(((((....((((((..........))))))......(((((((.((.....))(((.....((((((...))))))..)))...)))))))........).)))).... ( -32.10, z-score = -2.16, R) >droSim1.chr2R 14782311 113 - 19596830 UCAUGAUCAUCAUUUGGCACCGCCGCCCGCCAAACAAAAACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAG ...(((((....((((((..........))))))......(((((((.((.....))(((.....((((((...))))))..)))...)))))))........).)))).... ( -32.10, z-score = -2.16, R) >consensus UCAUGAUCAUCAUUUGGCACCGCCGCCCGCCAAACAAAAACUGCCACCGCCACCAGCGCCACCACAGCCACCCAGUGGCUCAGGCCCAGUGGCAGCACCACUAGAAUCAGCAG ...((((.....((((((..........))))))......(((((((.((.....))(((.....((((((...))))))..)))...)))))))..........)))).... (-31.50 = -31.50 + 0.00)


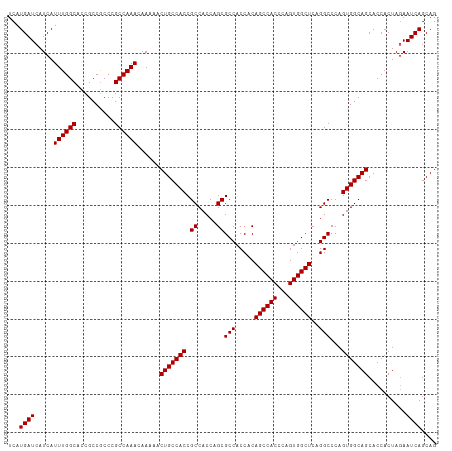
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:37:52 2011