
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 16,097,291 – 16,097,366 |
| Length | 75 |
| Max. P | 0.830589 |

| Location | 16,097,291 – 16,097,366 |
|---|---|
| Length | 75 |
| Sequences | 9 |
| Columns | 83 |
| Reading direction | forward |
| Mean pairwise identity | 76.86 |
| Shannon entropy | 0.46161 |
| G+C content | 0.55266 |
| Mean single sequence MFE | -25.77 |
| Consensus MFE | -16.19 |
| Energy contribution | -15.70 |
| Covariance contribution | -0.49 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.830589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 16097291 75 + 21146708 -----CGACC-CGGUGAUCAUUGUCGCAUCCGCCAC--GAUGGUGGCAAUUGUUUUGAUGGCCGCUGAUAACUGCAGCGGCGG -----....(-(.((.((((((((.((....)).))--)))))).)).............(((((((.......))))))))) ( -28.60, z-score = -1.33, R) >droSec1.super_1 13657946 75 + 14215200 -----CGACC-CGGUGAUCAUUGUCGCAUCCGCCAC--GAUGGUGGCAAUUGUUUUGAUGGCCGCUGAUAACUGCAGCGGCGG -----....(-(.((.((((((((.((....)).))--)))))).)).............(((((((.......))))))))) ( -28.60, z-score = -1.33, R) >droYak2.chr2R 8032113 75 - 21139217 -----CGACC-CGGUGAUCAUUGUCGCAUCCGCCAC--GAUGGUGGCAAUUGUUUUGAUGGCCGCUGAUAACUGCAGCGGCGG -----....(-(.((.((((((((.((....)).))--)))))).)).............(((((((.......))))))))) ( -28.60, z-score = -1.33, R) >droEre2.scaffold_4845 10247586 76 + 22589142 -----CGACCACGGCGAUCAUUGUCGCAUCCGCCAC--GAUGGUGGCAAUUGUUUUGAUGGCCGCUGAUAACUGCAACGGCGG -----......(((((..(((((..(((...(((((--....)))))...)))..)))))..)))))....((((....)))) ( -27.80, z-score = -1.35, R) >droAna3.scaffold_13266 10641074 80 - 19884421 -CCGCCAGUCUGGGCGAUCAUUGUCGCAUCCGCCAC--GAUGGUGGCAAUUGUUUUGAUGGCUGUUGAUAACUGCGAAGCGGG -((((((((.(.((((..(((((..(((...(((((--....)))))...)))..)))))..)))).)..))))....)))). ( -26.90, z-score = -0.45, R) >dp4.chr3 13233392 73 + 19779522 ----------UACCCGAUCAUUGUCGCAUCCGCCACCCGAUGGUGGCAAUUGUUUUGAUGGCUGUUGAUAACCGCAACAGCAG ----------........(((((..(((...((((((....))))))...)))..)))))(((((((.......))))))).. ( -25.90, z-score = -2.94, R) >droPer1.super_2 645804 73 - 9036312 ----------UACCCGAUCAUUGUCGCAUCCGCCACCCGAUGGUGGCAAUUGUUUUGAUGGCUGUUGAUAACCGCAACAGCAG ----------........(((((..(((...((((((....))))))...)))..)))))(((((((.......))))))).. ( -25.90, z-score = -2.94, R) >droWil1.scaffold_180701 2582292 71 + 3904529 ----------CGACGGGCCGUUGUCGUAUUCGCCAC--AGUGGUGGCAAUUGUUUUGAUGACUGUUGAUAACUGAAGCAGCAG ----------((((((....)))))).....(((((--....)))))..............((((((..........)))))) ( -17.70, z-score = 0.33, R) >droGri2.scaffold_15112 2754460 82 + 5172618 CGAAACGAAAACCGCCACCGCUUCCACUCCUUGCAUC-GAUGGUGGCAAUUGUUUUGAUGACCGUUGAUAACUGCAGCAGCGG ((((((((...((((((.((....((.....))...)-).))))))...))))))))....((((((..........)))))) ( -21.90, z-score = -1.02, R) >consensus _____CGACC_CGGCGAUCAUUGUCGCAUCCGCCAC__GAUGGUGGCAAUUGUUUUGAUGGCCGUUGAUAACUGCAGCGGCGG ..................(((((..(((...(((((......)))))...)))..))))).((((((..........)))))) (-16.19 = -15.70 + -0.49)


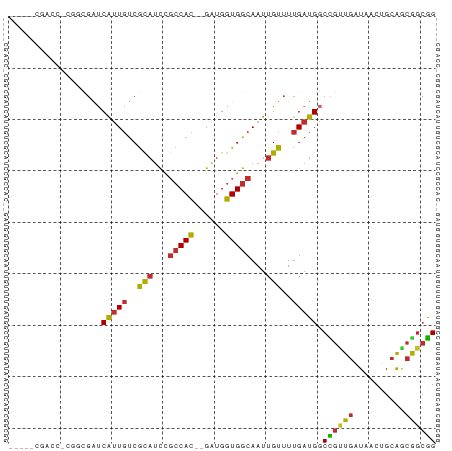
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:37:36 2011