
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 16,045,778 – 16,045,874 |
| Length | 96 |
| Max. P | 0.868713 |

| Location | 16,045,778 – 16,045,874 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 74.83 |
| Shannon entropy | 0.52730 |
| G+C content | 0.40777 |
| Mean single sequence MFE | -23.03 |
| Consensus MFE | -10.33 |
| Energy contribution | -10.53 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.868713 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
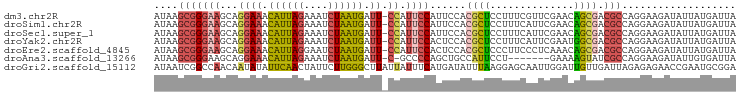
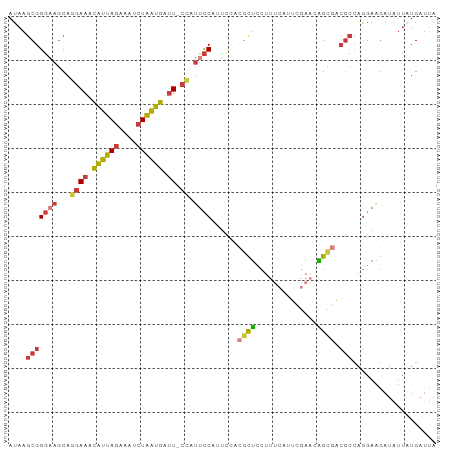
>dm3.chr2R 16045778 96 + 21146708 AUAAGCGGGAAGCAGGAAACAUUAGAAAUCUAAUGAUU-CCAUUCCAUUCCACGCUCCUUUCGUUCGAACAGCGACGCCAGGAAGAUAUUAUGAUUA ....(((((((...((((.((((((....)))))).))-))......)))).)))((((..(((.((.....)))))..)))).............. ( -26.90, z-score = -3.03, R) >droSim1.chr2R 14698631 96 + 19596830 AUAAGCGGGAAGCAGGAAACAUUAGAAAUCUAAUGAUU-CCAUUCCAUUCCACGCUCCUUUCAUUCGAACAGCGACGCCAGGAAGAUAUUAUGAUUA ....(((((((...((((.((((((....)))))).))-)).))))......((((...(((....))).)))).)))................... ( -24.10, z-score = -2.48, R) >droSec1.super_1 13607247 96 + 14215200 AUAAGCGGGAAGCAGGAAACAUUAGAAAUCUAAUGAUU-CCAUUCCAUUCCACGCUCCUUUCAUUCGAACAGCGACGCCAGGAAGAUAUUAUGAUUA ....(((((((...((((.((((((....)))))).))-)).))))......((((...(((....))).)))).)))................... ( -24.10, z-score = -2.48, R) >droYak2.chr2R 7979073 96 - 21139217 AUAAGCGGGAAGCAGGAAACAUUAGAAAUCUAAUGAUU-CCAUUCCACUCCACGCUCCUUUCAUUCGAAUGGCGACGCCAGGAAGAUAUUAUGAUUA ....(((((((...((((.((((((....)))))).))-)).))))......((((...(((....))).)))).)))................... ( -24.80, z-score = -2.01, R) >droEre2.scaffold_4845 10195608 96 + 22589142 AUAAGCGGGAAGCAGGAAACAUUAGGAAUCUAAUGAUU-CCAUUCCACUCCACGCUCCCUUCCCUCAAACAGCGACGCCAGGAAGAUAUUAUGAUUA ...(((((((....(....)....((((((....))))-)).......))).))))..(((((.................)))))............ ( -22.93, z-score = -1.59, R) >droAna3.scaffold_13266 10590474 88 - 19884421 AUAAGCGGGAAGCAGGAAACAUUAGAAAUCUAAUGAUU-C-GCCCCAGCUGCCAUUCCU-------GAAAAGUAUCGCCAGGAAGAUAUUGUGAUUA ...((((((..((..(((.((((((....)))))).))-)-))))).)))....(((((-------(...........))))))............. ( -21.50, z-score = -1.35, R) >droGri2.scaffold_15112 2487718 97 - 5172618 AUAAUCGGCCAACAAUAUAUUCAACUAUUCUUGGGCUUAUUAUUUCAUGAUAUUUAAGGAGCAAUUGGAUUGUUGAUUAGAGAGAACCGAAUGCGGA ....((((.........((.(((((.((((....((((..((((....))))......))))....)))).))))).)).......))))....... ( -16.89, z-score = 0.06, R) >consensus AUAAGCGGGAAGCAGGAAACAUUAGAAAUCUAAUGAUU_CCAUUCCAUUCCACGCUCCUUUCAUUCGAACAGCGACGCCAGGAAGAUAUUAUGAUUA ....(((((((...((.((((((((....)))))).)).)).))))......((((..............)))).)))................... (-10.33 = -10.53 + 0.19)

| Location | 16,045,778 – 16,045,874 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 74.83 |
| Shannon entropy | 0.52730 |
| G+C content | 0.40777 |
| Mean single sequence MFE | -24.87 |
| Consensus MFE | -11.36 |
| Energy contribution | -11.86 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.53 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.751573 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
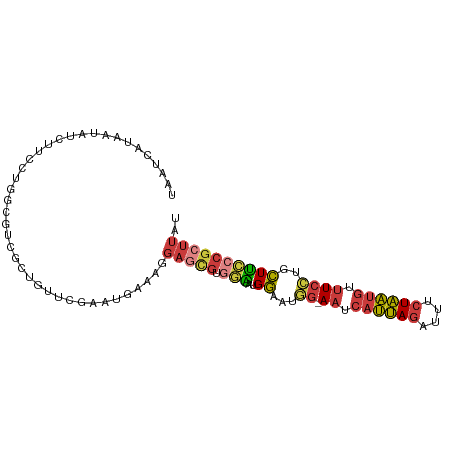
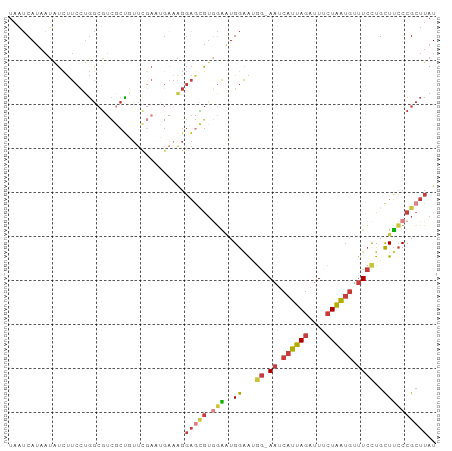
>dm3.chr2R 16045778 96 - 21146708 UAAUCAUAAUAUCUUCCUGGCGUCGCUGUUCGAACGAAAGGAGCGUGGAAUGGAAUGG-AAUCAUUAGAUUUCUAAUGUUUCCUGCUUCCCGCUUAU ..............((((..(((((.....)).)))..))))(((.(((..((...((-((.((((((....)))))).))))..)))))))).... ( -28.90, z-score = -2.64, R) >droSim1.chr2R 14698631 96 - 19596830 UAAUCAUAAUAUCUUCCUGGCGUCGCUGUUCGAAUGAAAGGAGCGUGGAAUGGAAUGG-AAUCAUUAGAUUUCUAAUGUUUCCUGCUUCCCGCUUAU ..............((((..(((((.....)).)))..))))(((.(((..((...((-((.((((((....)))))).))))..)))))))).... ( -27.10, z-score = -2.12, R) >droSec1.super_1 13607247 96 - 14215200 UAAUCAUAAUAUCUUCCUGGCGUCGCUGUUCGAAUGAAAGGAGCGUGGAAUGGAAUGG-AAUCAUUAGAUUUCUAAUGUUUCCUGCUUCCCGCUUAU ..............((((..(((((.....)).)))..))))(((.(((..((...((-((.((((((....)))))).))))..)))))))).... ( -27.10, z-score = -2.12, R) >droYak2.chr2R 7979073 96 + 21139217 UAAUCAUAAUAUCUUCCUGGCGUCGCCAUUCGAAUGAAAGGAGCGUGGAGUGGAAUGG-AAUCAUUAGAUUUCUAAUGUUUCCUGCUUCCCGCUUAU ..............((((..(((((.....)).)))..))))(((.(((..((...((-((.((((((....)))))).))))..)))))))).... ( -27.10, z-score = -1.78, R) >droEre2.scaffold_4845 10195608 96 - 22589142 UAAUCAUAAUAUCUUCCUGGCGUCGCUGUUUGAGGGAAGGGAGCGUGGAGUGGAAUGG-AAUCAUUAGAUUCCUAAUGUUUCCUGCUUCCCGCUUAU ...........(((((((..((........)).)))))))(((((.(((((((((.((-((((....))))))......)))).))))).))))).. ( -31.50, z-score = -2.06, R) >droAna3.scaffold_13266 10590474 88 + 19884421 UAAUCACAAUAUCUUCCUGGCGAUACUUUUC-------AGGAAUGGCAGCUGGGGC-G-AAUCAUUAGAUUUCUAAUGUUUCCUGCUUCCCGCUUAU .............(((((((.........))-------)))))....(((.(((((-(-((.((((((....)))))).)))..)).))).)))... ( -21.30, z-score = -1.05, R) >droGri2.scaffold_15112 2487718 97 + 5172618 UCCGCAUUCGGUUCUCUCUAAUCAACAAUCCAAUUGCUCCUUAAAUAUCAUGAAAUAAUAAGCCCAAGAAUAGUUGAAUAUAUUGUUGGCCGAUUAU ...(((...((..................))...))).................(((((..(((.((.((((........)))).)))))..))))) ( -11.07, z-score = 0.62, R) >consensus UAAUCAUAAUAUCUUCCUGGCGUCGCUGUUCGAAUGAAAGGAGCGUGGAAUGGAAUGG_AAUCAUUAGAUUUCUAAUGUUUCCUGCUUCCCGCUUAU ........................................(((((.((((.((((.......((((((....)))))).))))...))))))))).. (-11.36 = -11.86 + 0.50)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:37:21 2011