| Sequence ID | dm3.chr2R |
|---|---|
| Location | 15,632,646 – 15,632,742 |
| Length | 96 |
| Max. P | 0.981707 |

| Location | 15,632,646 – 15,632,742 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 72.44 |
| Shannon entropy | 0.37738 |
| G+C content | 0.50859 |
| Mean single sequence MFE | -28.13 |
| Consensus MFE | -24.99 |
| Energy contribution | -24.33 |
| Covariance contribution | -0.65 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.89 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.08 |
| SVM RNA-class probability | 0.981707 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 15632646 96 + 21146708 CCCUCAUCUAAGUACUAACCGCGCCCGACGCUGCUUAAUUUCGGUGAUCGGACGAGAACCGAUGUAUUCAGCGUGGUAUGGUCGUUGGCGAAAGUC .................((((..((..((((((..((...(((((..((....))..)))))..))..))))))))..))))....(((....))) ( -31.10, z-score = -1.32, R) >anoGam1.chr2L 13376025 91 + 48795086 CACCCACCUAAGUACUAGCCAUGCCCGAUGUUGCUUAACUUCGGUGAUCGGACGAGAACCGGUGUUUUCAACAUGGUAUGGUCGUUGACAA----- ...........((....((((((((..((((((...(((.(((((..((....))..))))).)))..)))))))))))))).....))..----- ( -29.20, z-score = -2.02, R) >apiMel3.GroupUn 29499569 86 + 399230636 -AGCCAUCCAAAUAUUAACCGUGCCCGACGCAGCUUGACUUUGCUGAUCGAACGAGAACCAAUAGUUCCUACGUGGUACGGUCGUUG--------- -................(((((((((((..((((........)))).))).(((.((((.....))))...))))))))))).....--------- ( -24.10, z-score = -2.11, R) >consensus CACCCAUCUAAGUACUAACCGUGCCCGACGCUGCUUAACUUCGGUGAUCGGACGAGAACCGAUGUUUUCAACGUGGUAUGGUCGUUG_C_A_____ .................((((((((..((((((...(((.(((((..((....))..))))).)))..)))))))))))))).............. (-24.99 = -24.33 + -0.65)
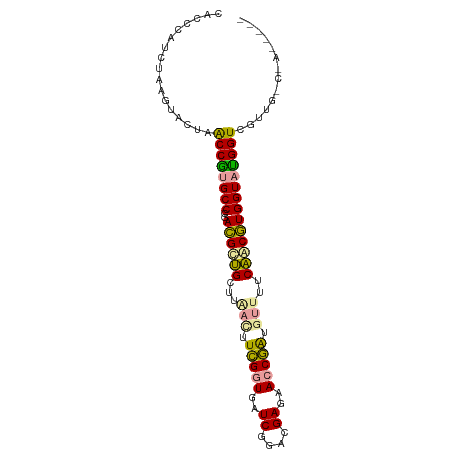



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:35:52 2011