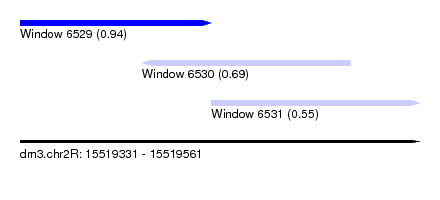

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 15,519,331 – 15,519,561 |
| Length | 230 |
| Max. P | 0.939254 |

| Location | 15,519,331 – 15,519,441 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 87.68 |
| Shannon entropy | 0.16977 |
| G+C content | 0.40201 |
| Mean single sequence MFE | -30.15 |
| Consensus MFE | -24.46 |
| Energy contribution | -25.14 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.43 |
| SVM RNA-class probability | 0.939254 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
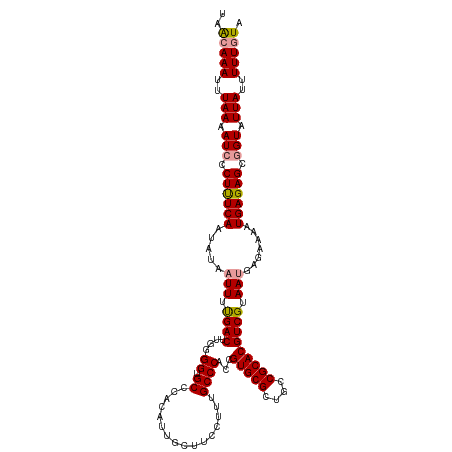
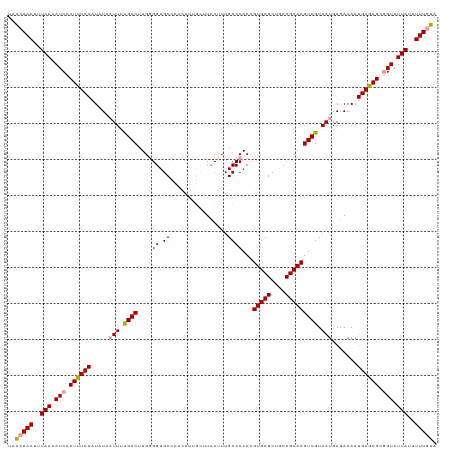
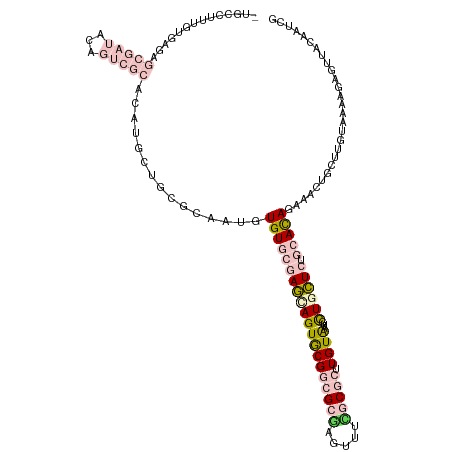
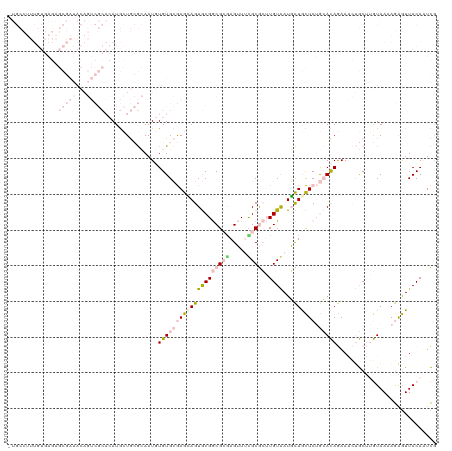
>dm3.chr2R 15519331 110 + 21146708 UAGAAAAUUUAAGAUCCCUCUCAAUAUAUUUACGACUUGUGGUGC---------UUUCAUUGCCCACCGUGCGCUACCGCACGUCGUAAUGAGAAAAUGAGAGCUGUAUUAUUUUUGUA ..((((((.....((..((((((......(((((((..((((.((---------.......)))))).(((((....))))))))))))........))))))..))...))))))... ( -31.94, z-score = -4.25, R) >droSec1.super_1 13034014 119 + 14215200 UAACAAAUUUAAAAUCCCUUUCAAUAUAAUUUUGACUUGGGGUGCCCACAUUGCUUCCUUUGCCCACAGUGCGCUGCCGCACGUCGUAAUGAGAAAAUGAGAGCGGUAUUAUUUUUGUA ..(((((..(((.(((.((((((.....(((.((((.((.(((((.(((...((.......)).....))).)).))).)).)))).))).......)))))).))).)))..))))). ( -29.20, z-score = -1.39, R) >droSim1.chr2R 14223085 119 + 19596830 UAACAAAUUUAAAAUCCCUUUCAAUAUAAUUUUGACUUGGGGUGCCCACAUUGCUUCCUUUGCCCACCGUGCGCUGCCGCACGUCGUAAUGAGAAAAUGAGAGCGGUAUUAUUUUUGUA ..(((((..(((.(((((..((((.......))))...)))))((.(.((((..(((..((((....((((((....))))))..)))).)))..)))).).))....)))..))))). ( -29.30, z-score = -1.49, R) >consensus UAACAAAUUUAAAAUCCCUUUCAAUAUAAUUUUGACUUGGGGUGCCCACAUUGCUUCCUUUGCCCACCGUGCGCUGCCGCACGUCGUAAUGAGAAAAUGAGAGCGGUAUUAUUUUUGUA ..(((((..(((.(((.((((((.....(((.((((...((((..................))))...(((((....))))))))).))).......)))))).))).)))..))))). (-24.46 = -25.14 + 0.67)

| Location | 15,519,401 – 15,519,521 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.83 |
| Shannon entropy | 0.10980 |
| G+C content | 0.47742 |
| Mean single sequence MFE | -27.80 |
| Consensus MFE | -26.14 |
| Energy contribution | -25.98 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.94 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.689773 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
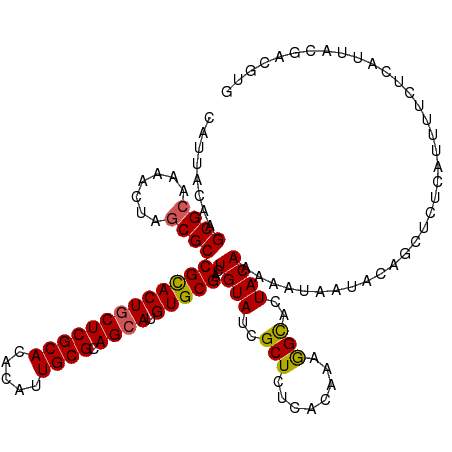
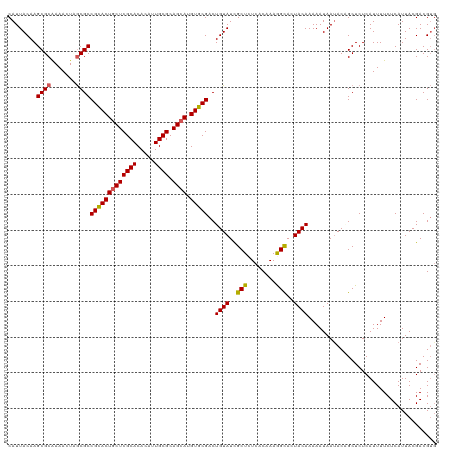
>dm3.chr2R 15519401 120 - 21146708 CAUUACAAGCGCAAAACUCGCGCCGCACUACUCGCACACAUUGCGCAGCAUGUGCGACUGUAUCGCUCUCACAAAGGCAGUACAAAAAUAAUACAGCUCUCAUUUUCUCAUUACGACGUG ........((((.......))))(((((..((((((.....)))).))...)))))..(((((.(((........))).))))).................................... ( -25.30, z-score = -0.36, R) >droEre2.scaffold_4845 9717831 119 - 22589142 CAUUACAAGCGCAAAACUAACGCCGUACUGCUCGCACACACUGCGCAGCAUGUGCGACUGUAUUGCUCUCACUAAAGCACUACAAAAAUAAUACAGCUCUCAUUUUCUCAUUACGACGG- .......(((.............(((((((((((((.....)))).)))).)))))..((((.((((........)))).))))...........))).....................- ( -25.70, z-score = -1.57, R) >droYak2.chr2R 7484240 120 + 21139217 CAUUACAAGCGCAAAACUAGCGCCGCACUGCUCGCACACACUGCGCAGCAUGUGCGACUGUAUCGCUCUCACUAAAGUACUACAAAAAUAACACGGCUCUCAUUUUCUCAUUACGACGUG ........((((.......))))(((((((((((((.....)))).)))).)))))..((((..(((........)))..)))).......((((.....................)))) ( -29.30, z-score = -1.66, R) >droSec1.super_1 13034093 120 - 14215200 CAUUACAAGCGCGAAACUUGCGCCGCACUGCUCGCACACAUUGCGCAGCAUGUGCGACUGUAUCGCUCUCACAAAGGCACUACAAAAAUAAUACCGCUCUCAUUUUCUCAUUACGACGUG ........((((((...))))))(((((((((((((.....)))).)))).)))))..((((..(((........)))..)))).................................... ( -29.20, z-score = -1.08, R) >droSim1.chr2R 14223164 120 - 19596830 CAUUACAAGCGCAAAACUUGCGCCGCACUGCUCGCACACAUUGCGCAGCAUGUGCGACUGUAUCGCUCUCACAAAGGCACUACAAAAAUAAUACCGCUCUCAUUUUCUCAUUACGACGUG ........(((((.....)))))(((((((((((((.....)))).)))).)))))..((((..(((........)))..)))).................................... ( -29.50, z-score = -1.59, R) >consensus CAUUACAAGCGCAAAACUAGCGCCGCACUGCUCGCACACAUUGCGCAGCAUGUGCGACUGUAUCGCUCUCACAAAGGCACUACAAAAAUAAUACAGCUCUCAUUUUCUCAUUACGACGUG ........((((.......))))(((((((((((((.....)))).)))).)))))..((((..(((........)))..)))).................................... (-26.14 = -25.98 + -0.16)

| Location | 15,519,441 – 15,519,561 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 62.25 |
| Shannon entropy | 0.59984 |
| G+C content | 0.41110 |
| Mean single sequence MFE | -33.30 |
| Consensus MFE | -10.98 |
| Energy contribution | -12.58 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.551407 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 15519441 120 + 21146708 CUGCCUUUGUGAGAGCGAUACAGUCGCACAUGCUGCGCAAUGUGUGCGAGUAGUGCGGCGCGAGUUUUGCGCUUGUAAUGCUGCUCUGCACAGAAACUGCUGGUAAAAGAGUUACAAUCG .....((((((((((((.....(((((((.((((((((.....)))).)))))))))))((((((.....)))))).....)))))).))))))((((.((......))))))....... ( -43.40, z-score = -0.50, R) >droSec1.super_1 13034133 120 + 14215200 GUGCCUUUGUGAGAGCGAUACAGUCGCACAUGCUGCGCAAUGUGUGCGAGCAGUGCGGCGCAAGUUUCGCGCUUGUAAUGCUGCUCUGCACAGAAACUGCUUGUAAAAGAGUUACAUUCG .....((((((((((((.....(((((((.((((((((.....)))).)))))))))))((((((.....)))))).....)))))).))))))((((.(((....)))))))....... ( -46.40, z-score = -0.97, R) >droSim1.chr2R 14223204 120 + 19596830 GUGCCUUUGUGAGAGCGAUACAGUCGCACAUGCUGCGCAAUGUGUGCGAGCAGUGCGGCGCAAGUUUUGCGCUUGUAAUGCUGCUCUGCACAGAAACUGCUGGUAAAAGAGUUACAUUCG .....((((((((((((.....(((((((.((((((((.....)))).)))))))))))((((((.....)))))).....)))))).))))))((((.((......))))))....... ( -45.70, z-score = -0.69, R) >dp4.chrXR_group6 7130190 91 - 13314419 ----------------------------AAUAAUAAAGAAU-UGUGGAAAGAGUACGACGAGAGAUGCGCAAAUGUGAUUUUAUUCAAAAUAAAAUACACAUAUAAAAAGAUUACAAUUA ----------------------------..........(((-(((((.....(..((.(....).))..)..((((((((((((.....))))))).))))).........)))))))). ( -15.50, z-score = -3.24, R) >droPer1.super_36 61845 91 + 818889 ----------------------------AAUAAUAAAGAAU-UGUGGAAAGAGUACGACGAGAGAUGCGCAAAUGUGAUUUUAUUCAAAAUAAAAUACACAUAUAAAAAGAUUACAAUUA ----------------------------..........(((-(((((.....(..((.(....).))..)..((((((((((((.....))))))).))))).........)))))))). ( -15.50, z-score = -3.24, R) >consensus _UGCCUUUGUGAGAGCGAUACAGUCGCACAUGCUGCGCAAUGUGUGCGAGCAGUGCGGCGCGAGUUUCGCGCUUGUAAUGCUGCUCUGCACAGAAACUGCUUGUAAAAGAGUUACAAUCG ..............((((.....))))...............((((((((((((((((((((.....))))).))))...)))))).)))))............................ (-10.98 = -12.58 + 1.60)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:35:23 2011