
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 15,059,017 – 15,059,120 |
| Length | 103 |
| Max. P | 0.578831 |

| Location | 15,059,017 – 15,059,120 |
|---|---|
| Length | 103 |
| Sequences | 8 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 67.35 |
| Shannon entropy | 0.69748 |
| G+C content | 0.54987 |
| Mean single sequence MFE | -33.48 |
| Consensus MFE | -11.66 |
| Energy contribution | -12.90 |
| Covariance contribution | 1.24 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.578831 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 15059017 103 - 21146708 GUUACGGCUGAGCGAGUUCCCAGUUUGCCAGUUGGACGGCGAAGGGGAAAGAGCUGGAGCCGAAGCAGCU-GAACAUCGAUUUCAAGUUGAAUGUCGUGGCGGG (((((((((.(((...(((((..((((((........)))))).)))))...))).).)))((..(((((-(((.......))).)))))....)))))))... ( -38.30, z-score = -1.97, R) >droSec1.super_1 12571805 103 - 14215200 GUUACGGCUGAGCGAGUUGCCAGUUCGCCAGUUGGACGGCGAAAGGGAAAGAGCUGGAGCCGAAGCAGCU-GAACAUCGAUUUCAAGUUGAAUGUCGUGGCGGG ((((((((((..((.(((.(((((((.((..((.(....).))..))...))))))))))))...)))).-..((((((((.....)))).))))))))))... ( -36.00, z-score = -1.23, R) >droSim1.chr2R 13763342 103 - 19596830 GUUACGGCUGAGCGAGUUCCCAGUUCGCCAGUUGGCCGGCGAAGGGGAAAGAGCUGGAGCCGAAGCAGCU-GAACAUCGAUUUCAAGUUGAAUGUCGUGGCGGG (((((((((.(((...(((((..((((((.(....).)))))).)))))...))).).)))((..(((((-(((.......))).)))))....)))))))... ( -40.70, z-score = -2.02, R) >droYak2.chr2R 7018897 91 + 21139217 GUUACGGCUGAGCGAGUUCCCAGUUCGCCGGCUGGCUGGAGAAGGGGAAA------------AAGCAGCU-GAACAUCGAUUUCAAGUUGAAUGUCGUGGCGGG ((((((((...((...(((((..(((.((((....)))).))).))))).------------..))....-.....(((((.....)))))..))))))))... ( -33.10, z-score = -1.59, R) >droEre2.scaffold_4845 9259152 103 - 22589142 GUUACGGCUGAGCGAGUUCCCAGUUCGCCAGUUGGCUGGAGGAGGGGAAAAGCGCUGGGGCCAAGCAGCU-GAACAUCGAUUUCAAGUUGAAUGUCGUGGCGGG (((((((((.((((..(((((..(((.((((....)))).))).)))))...)))).).)))...(((((-(((.......))).)))))......)))))... ( -39.60, z-score = -1.60, R) >droAna3.scaffold_13266 8384312 95 + 19884421 GUUACGGCUGAGCGAGUUCCCAGCUUG-GAGCUGCGAGGGGCUCGGAGCAGCGG--------AGGCAGCUCUAACAUCGAUUUCAAGUUGAAUGUCGUGGCGGG ((((((((.((((..((..((.((((.-(((((......))))).))))...))--------..)).)))).....(((((.....)))))..))))))))... ( -37.90, z-score = -1.36, R) >droWil1.scaffold_181009 1755288 86 - 3585778 GUUUCGCUUGAGCGAAUGCCU-GCCUGCCAUAUGGGAAAAACAUGAGAGAGAGA---ACAUGGAGAAGCC-UAA-AUCGAUUUCAAGUUGAA------------ ..(((((....))))).....-.(((.......)))...(((.(((((..((..---....((.....))-...-.))..))))).)))...------------ ( -15.10, z-score = 0.83, R) >droGri2.scaffold_15112 403278 93 + 5172618 GCCAUGGCUGAGUUGCUGAGCCGAUGAGCAACUGUCUACCCAAUUAAGUCUUGA--------UGGUAGUA-AAAGGCCG--CUCAAGUUGAAAGUCGAGGCCAC (((.(((((.(((((((.(.....).)))))))................(((((--------((((....-....))))--.))))).....))))).)))... ( -27.10, z-score = -1.10, R) >consensus GUUACGGCUGAGCGAGUUCCCAGUUCGCCAGUUGGCCGGCGAAGGGGAAAGAGC___AG_CGAAGCAGCU_GAACAUCGAUUUCAAGUUGAAUGUCGUGGCGGG ....((((((.(((((.......))))))))))).................................(((...((((((((.....)))).))))...)))... (-11.66 = -12.90 + 1.24)


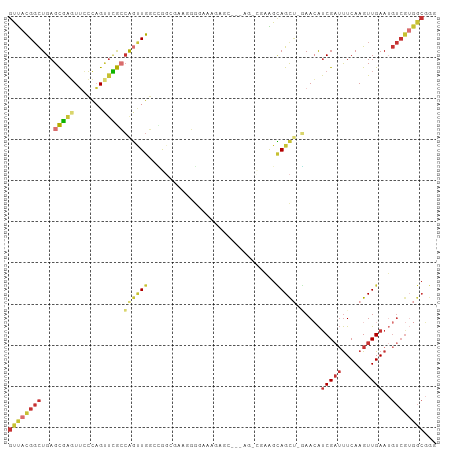
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:34:08 2011