
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 14,995,095 – 14,995,212 |
| Length | 117 |
| Max. P | 0.734601 |

| Location | 14,995,095 – 14,995,212 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 63.96 |
| Shannon entropy | 0.59165 |
| G+C content | 0.35791 |
| Mean single sequence MFE | -24.93 |
| Consensus MFE | -13.64 |
| Energy contribution | -13.45 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.58 |
| Mean z-score | -0.77 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.657072 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
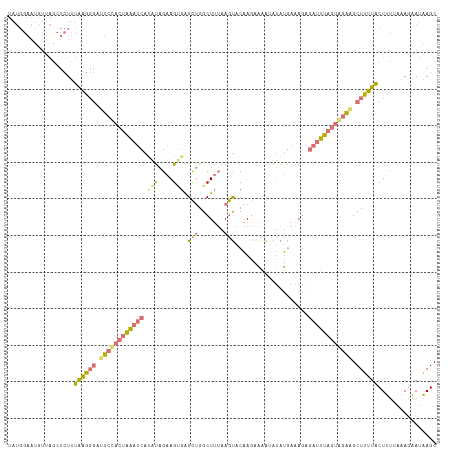
>dm3.chr2R 14995095 117 + 21146708 UAUGGAAUAUUAGCUCUUUAAGGGAUUCUACUAAAUCACGUAGAAGCGAGCUGGCUUUAAGUACAAGAAGAGAUAUAAAAGAGAUUUAGUAGAAGCUUUUACUUUAAAAUAAUAAGC .......(((((......((((((.((((((((((((((...(((((......)))))..)).(.....)............)))))))))))).))))))........)))))... ( -22.44, z-score = -0.50, R) >droSim1.chr2R 13696543 117 + 19596830 UAUGGAAUGUUAGCUCUUUAAGGGAUUCCACUAAAUCAUAUAGAAGUGAGCUGGCUUUAAGUACAAGAAAAUAUAUGAAAGAGAUUUAGUGGAAGCUUUUACUUUUAAAGAAUAAGC .........(((..((((((((((.(((((((((((((((((....((.(((.......))).))......)))))))......)))))))))).......)))))))))).))).. ( -25.80, z-score = -1.73, R) >droSec1.super_1 12508063 117 + 14215200 UAUGGAAUGUUAGCUCUUUAAGGGAGUCCACUAAAUCAUAUAGAAGUGAGCUGGCUUUAAGUACAAGAAAAUAUAUGAAAGAGAUUUAGUGGAAGCUUUUACUUUUAAAGAAUAAGC .........(((..((((((((((..((((((((((((((((....((.(((.......))).))......)))))))......)))))))))........)))))))))).))).. ( -24.70, z-score = -1.22, R) >droVir3.scaffold_12875 1174408 96 - 20611582 ----------------UGUGGUUAACAACCUUUGAACUGACAGCAG--GGUCGGCUAAAAACACUACCCAGCACUGGCUAAAAGCUGCUUAAUGGCUGCUGCUGC---AGCAUCAGC ----------------...(((.....)))......((((((((.(--(((.(..........).))))(((....)))....)))).......(((((....))---))).)))). ( -26.80, z-score = 0.37, R) >consensus UAUGGAAUGUUAGCUCUUUAAGGGAUUCCACUAAAUCAUAUAGAAGUGAGCUGGCUUUAAGUACAAGAAAAUAUAUGAAAGAGAUUUAGUAGAAGCUUUUACUUUUAAAGAAUAAGC ..................((((((.((((((((((((.(((....))).(((.......)))....................)))))))))))).))))))................ (-13.64 = -13.45 + -0.19)


| Location | 14,995,095 – 14,995,212 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 63.96 |
| Shannon entropy | 0.59165 |
| G+C content | 0.35791 |
| Mean single sequence MFE | -21.43 |
| Consensus MFE | -9.84 |
| Energy contribution | -10.28 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.734601 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
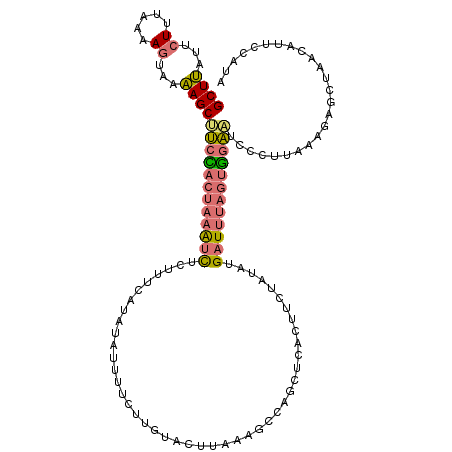
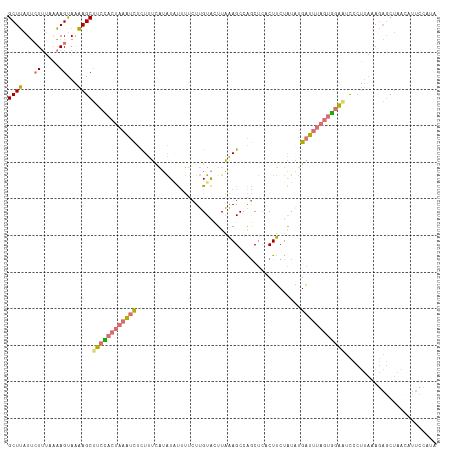
>dm3.chr2R 14995095 117 - 21146708 GCUUAUUAUUUUAAAGUAAAAGCUUCUACUAAAUCUCUUUUAUAUCUCUUCUUGUACUUAAAGCCAGCUCGCUUCUACGUGAUUUAGUAGAAUCCCUUAAAGAGCUAAUAUUCCAUA ((((.(((((....)))))))))((((((((((((.(...((((........))))....((((......))))....).))))))))))))......................... ( -19.80, z-score = -1.65, R) >droSim1.chr2R 13696543 117 - 19596830 GCUUAUUCUUUAAAAGUAAAAGCUUCCACUAAAUCUCUUUCAUAUAUUUUCUUGUACUUAAAGCCAGCUCACUUCUAUAUGAUUUAGUGGAAUCCCUUAAAGAGCUAACAUUCCAUA .....((((((((..((....))((((((((((((.......((((......)))).....((....))...........))))))))))))....))))))))............. ( -18.90, z-score = -1.91, R) >droSec1.super_1 12508063 117 - 14215200 GCUUAUUCUUUAAAAGUAAAAGCUUCCACUAAAUCUCUUUCAUAUAUUUUCUUGUACUUAAAGCCAGCUCACUUCUAUAUGAUUUAGUGGACUCCCUUAAAGAGCUAACAUUCCAUA .....((((((((.(((....)))(((((((((((.......((((......)))).....((....))...........))))))))))).....))))))))............. ( -18.60, z-score = -1.81, R) >droVir3.scaffold_12875 1174408 96 + 20611582 GCUGAUGCU---GCAGCAGCAGCCAUUAAGCAGCUUUUAGCCAGUGCUGGGUAGUGUUUUUAGCCGACC--CUGCUGUCAGUUCAAAGGUUGUUAACCACA---------------- ..(((((((---((....)))).)))))(((((((((((((((((...((((.(.((.....))).)))--).))))...))).)))))))))).......---------------- ( -28.40, z-score = 0.35, R) >consensus GCUUAUUCUUUAAAAGUAAAAGCUUCCACUAAAUCUCUUUCAUAUAUUUUCUUGUACUUAAAGCCAGCUCACUUCUAUAUGAUUUAGUGGAAUCCCUUAAAGAGCUAACAUUCCAUA ((((...((.....))...))))((((((((((((.............................................))))))))))))......................... ( -9.84 = -10.28 + 0.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:33:57 2011