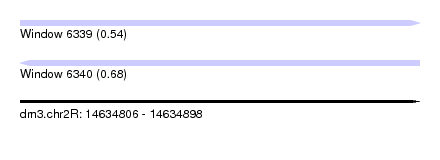
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 14,634,806 – 14,634,898 |
| Length | 92 |
| Max. P | 0.682460 |

| Location | 14,634,806 – 14,634,898 |
|---|---|
| Length | 92 |
| Sequences | 7 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 90.78 |
| Shannon entropy | 0.17880 |
| G+C content | 0.60409 |
| Mean single sequence MFE | -35.66 |
| Consensus MFE | -28.10 |
| Energy contribution | -29.71 |
| Covariance contribution | 1.61 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.535445 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
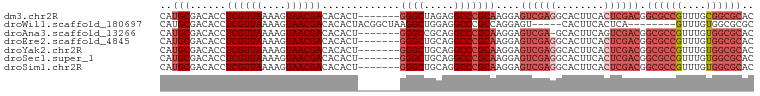
>dm3.chr2R 14634806 92 + 21146708 GUGCGCCGCAAACGGCGCCGUCGAGUGAAGUGCCUCGACUCCUUGCGGGCCUCUAGCCC-------AGUGUGUCGUUACUUUUAACGAGGUGUCGCAUG (.((((((....)))))))((((((........))))))....(((((((.....))))-------.(..(.((((((....)))))).)..).))).. ( -36.80, z-score = -2.06, R) >droWil1.scaffold_180697 2103255 86 + 4168966 GCGCGCCACAAAC--------UGAGUGAAGUG-----ACUCCUGGCGGGCCUCCAGCCUUAGCCGUAGUGUGUCGUUACUUUUAACGAGGUGUCGCAUG (((((((.((.((--------(((((......-----))))..(((((((.....))))..)))..))).))((((((....)))))))))).)))... ( -29.60, z-score = -1.15, R) >droAna3.scaffold_13266 2153059 91 - 19884421 GUGCGCCACAAACGGCGCCGUCGACUGAAGUGC-UCGACUCCUUGCGGGCCUGCGGCCC-------AGUGUGUCGUUACUUUUAACGAGGUGUCGCAUG (((((.(((.(((((((((((((((......).-))))).......(((((...)))))-------.).)))))))).(((.....)))))).))))). ( -36.40, z-score = -1.42, R) >droEre2.scaffold_4845 8826428 92 + 22589142 GUGCGCCACAAACGGCGCCGUCGAGUGAAGUGCCUCGACUCCUUGCGGGCCUGCAGCCC-------AGUGUGUCGUUACUUUUAACGAGGUGUCGCAUG (((((.(((.(((((((((((((((........)))))).......((((.....))))-------.).)))))))).(((.....)))))).))))). ( -36.70, z-score = -1.77, R) >droYak2.chr2R 6575970 92 - 21139217 GUGCGCCACAAACGGCGCCGUCGAGUGAAGUGCCUCGACUCCUUGCGGGCCUGCAGCCC-------AGUGUGUCGUUACUUUUAACGAGGUGUCGCAUG (((((.(((.(((((((((((((((........)))))).......((((.....))))-------.).)))))))).(((.....)))))).))))). ( -36.70, z-score = -1.77, R) >droSec1.super_1 12143466 92 + 14215200 GUGCGCCACAAACGGCGCCGUCGAGUGAAGUGCCUCGACUCCUUGCGGGCCUGCAGCCC-------AGUGUGUCGUUACUUUUAACGAGGUGUCGCAUG (((((.(((.(((((((((((((((........)))))).......((((.....))))-------.).)))))))).(((.....)))))).))))). ( -36.70, z-score = -1.77, R) >droSim1.chr2R 13351293 92 + 19596830 GUGCGCCACAAACGGCGCCGUCGAGUGAAGUGCCUCGACUCCUUGCGGGCCUGCAGCCC-------AGUGUGUCGUUACUUUUAACGAGGUGUCGCAUG (((((.(((.(((((((((((((((........)))))).......((((.....))))-------.).)))))))).(((.....)))))).))))). ( -36.70, z-score = -1.77, R) >consensus GUGCGCCACAAACGGCGCCGUCGAGUGAAGUGCCUCGACUCCUUGCGGGCCUGCAGCCC_______AGUGUGUCGUUACUUUUAACGAGGUGUCGCAUG (((((.(((.((((((((.((((((........)))))).......((((.....))))..........)))))))).(((.....)))))).))))). (-28.10 = -29.71 + 1.61)
| Location | 14,634,806 – 14,634,898 |
|---|---|
| Length | 92 |
| Sequences | 7 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 90.78 |
| Shannon entropy | 0.17880 |
| G+C content | 0.60409 |
| Mean single sequence MFE | -34.41 |
| Consensus MFE | -28.16 |
| Energy contribution | -30.04 |
| Covariance contribution | 1.88 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.682460 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
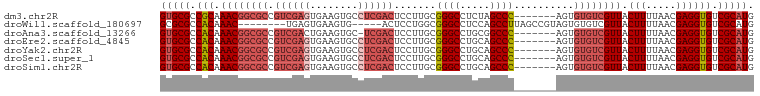
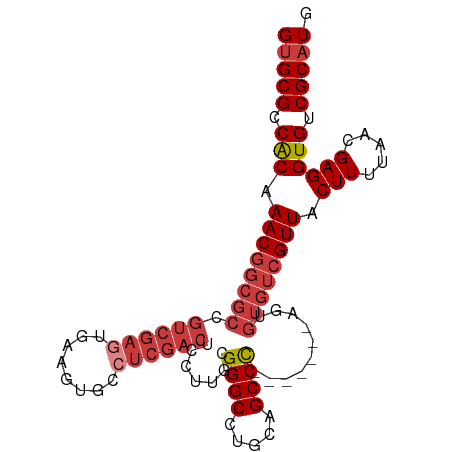
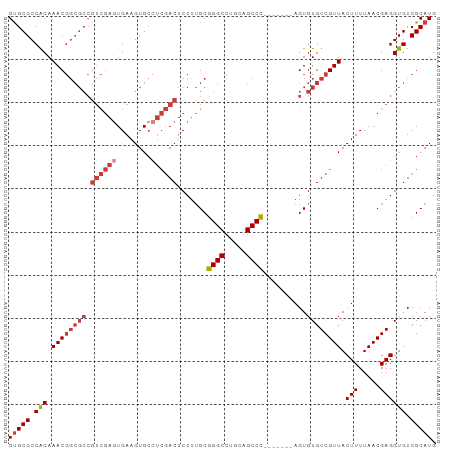
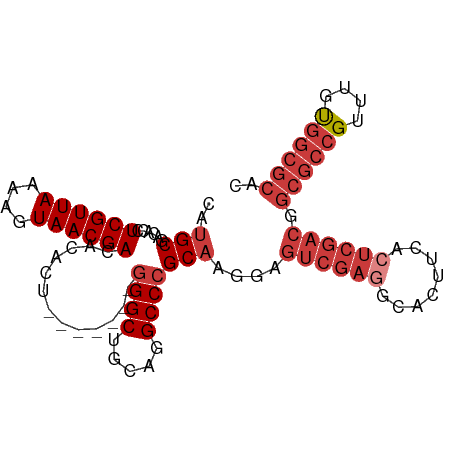
>dm3.chr2R 14634806 92 - 21146708 CAUGCGACACCUCGUUAAAAGUAACGACACACU-------GGGCUAGAGGCCCGCAAGGAGUCGAGGCACUUCACUCGACGGCGCCGUUUGCGGCGCAC ..(((......((((((....))))))......-------((((.....)))))))....((((((........)))))).((((((....)))))).. ( -38.80, z-score = -3.21, R) >droWil1.scaffold_180697 2103255 86 - 4168966 CAUGCGACACCUCGUUAAAAGUAACGACACACUACGGCUAAGGCUGGAGGCCCGCCAGGAGU-----CACUUCACUCA--------GUUUGUGGCGCGC ...(((.(...((((((....))))))((((..(((((...(((.....))).)))..((((-----......)))).--------)).)))))))).. ( -25.10, z-score = -0.20, R) >droAna3.scaffold_13266 2153059 91 + 19884421 CAUGCGACACCUCGUUAAAAGUAACGACACACU-------GGGCCGCAGGCCCGCAAGGAGUCGA-GCACUUCAGUCGACGGCGCCGUUUGUGGCGCAC ..(((......((((((....))))))......-------(((((...))))))))....(((((-.(......)))))).((((((....)))))).. ( -33.00, z-score = -0.95, R) >droEre2.scaffold_4845 8826428 92 - 22589142 CAUGCGACACCUCGUUAAAAGUAACGACACACU-------GGGCUGCAGGCCCGCAAGGAGUCGAGGCACUUCACUCGACGGCGCCGUUUGUGGCGCAC ..(((......((((((....))))))......-------((((.....)))))))....((((((........)))))).((((((....)))))).. ( -36.00, z-score = -1.89, R) >droYak2.chr2R 6575970 92 + 21139217 CAUGCGACACCUCGUUAAAAGUAACGACACACU-------GGGCUGCAGGCCCGCAAGGAGUCGAGGCACUUCACUCGACGGCGCCGUUUGUGGCGCAC ..(((......((((((....))))))......-------((((.....)))))))....((((((........)))))).((((((....)))))).. ( -36.00, z-score = -1.89, R) >droSec1.super_1 12143466 92 - 14215200 CAUGCGACACCUCGUUAAAAGUAACGACACACU-------GGGCUGCAGGCCCGCAAGGAGUCGAGGCACUUCACUCGACGGCGCCGUUUGUGGCGCAC ..(((......((((((....))))))......-------((((.....)))))))....((((((........)))))).((((((....)))))).. ( -36.00, z-score = -1.89, R) >droSim1.chr2R 13351293 92 - 19596830 CAUGCGACACCUCGUUAAAAGUAACGACACACU-------GGGCUGCAGGCCCGCAAGGAGUCGAGGCACUUCACUCGACGGCGCCGUUUGUGGCGCAC ..(((......((((((....))))))......-------((((.....)))))))....((((((........)))))).((((((....)))))).. ( -36.00, z-score = -1.89, R) >consensus CAUGCGACACCUCGUUAAAAGUAACGACACACU_______GGGCUGCAGGCCCGCAAGGAGUCGAGGCACUUCACUCGACGGCGCCGUUUGUGGCGCAC ..(((......((((((....)))))).............((((.....)))))))....((((((........)))))).((((((....)))))).. (-28.16 = -30.04 + 1.88)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:32:46 2011