
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 14,527,061 – 14,527,150 |
| Length | 89 |
| Max. P | 0.913879 |

| Location | 14,527,061 – 14,527,150 |
|---|---|
| Length | 89 |
| Sequences | 11 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 80.00 |
| Shannon entropy | 0.44020 |
| G+C content | 0.52276 |
| Mean single sequence MFE | -28.17 |
| Consensus MFE | -28.29 |
| Energy contribution | -27.15 |
| Covariance contribution | -1.13 |
| Combinations/Pair | 1.36 |
| Mean z-score | -0.85 |
| Structure conservation index | 1.00 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.913879 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 14527061 89 + 21146708 GACUCCGUGGCGCAACGG-UAGCGCGUCCGACUCCAGAUCGGAAGGUUGCGUGUUCAAAUCACGUCGGGGUCAA-GCAAUUGAUGUUUUUU------ (((((((..((((.....-..)))).((((((....).))))).....(((((.......))))))))))))((-(((.....)))))...------ ( -28.60, z-score = -0.64, R) >anoGam1.chr2R 5380048 96 - 62725911 GACUCCGUGGCGCAACGG-UAGCGCGUCUGACUCCAGAUCAGAAGGUUGCGUGUUCAAAUCACGUCGGGGUCAAAAAAUAUGAUGUGCAAUUCCUUU ....((((......))))-..((((((((((((((.((((....))))(((((.......))))).)))))))........)))))))......... ( -30.00, z-score = -1.51, R) >droGri2.scaffold_15245 13598005 90 + 18325388 GACUCCGUGGCGCAACGG-UAGCGCGUCUGACUCCAGAUCAGAAGGUUGCGUGUUCGAUUCACGUCGGGGUCAAG-AGCUCGAGUGAUUUUU----- .((((.((.((((.....-..)))).(((((((((.((((....))))(((((.......))))).)))))).))-)))..)))).......----- ( -27.70, z-score = 0.35, R) >droMoj3.scaffold_6496 15789347 88 + 26866924 GACUCCGUGGCGCAACGG-UAGCGCGUCUGACUCCAGAUCAGAAGGUUGCGUGUUCGAUUCACGUCGGGGUCA---AAUGUGACAAAUUAUU----- ..((((((......))))-.))(((((.(((((((.((((....))))(((((.......))))).)))))))---.)))))..........----- ( -28.00, z-score = -0.91, R) >droVir3.scaffold_12875 9473643 84 - 20611582 GACUCCGUGGCGCAACGG-UAGCGCGUCUGACUCCAGAUCAGAAGGUUGCGUGUUCGAUUCACGUCGGGGUCAAUCGAGGUUUUU------------ ..(((....((((.....-..))))(..(((((((.((((....))))(((((.......))))).)))))))..))))......------------ ( -26.50, z-score = -0.15, R) >droWil1.scaffold_180700 1819304 88 - 6630534 GACUCCGUGGCGCAACGG-UAGCGCGUCUGACUCCAGAUCAGAAGGUUGCGUGUUCAAAUCACGUCGGGGUCAACGAUUGGGAUUUCUU-------- ...(((...((((.....-..))))((.(((((((.((((....))))(((((.......))))).))))))))).....)))......-------- ( -27.00, z-score = -0.20, R) >droPer1.super_31 71934 91 + 935084 GACUCCGUGGCGCAACGG-UAGCGCGUCUGACUCCAGAUCAGAAGGUUGCGUGUUCAAAUCACGUCGGGGUCAAUAGACUAAUUUUUUUAAA----- (((((((..((((.....-..)))).((((((....).))))).....(((((.......))))))))))))....................----- ( -26.40, z-score = -1.28, R) >droAna3.scaffold_13266 17227175 92 - 19884421 GGUCCUGUGGCGCAAUGGAUAACGCGUCUGACUACGGAUCAGAAGAUUCCAGGUUCGACUCCUGGCAGGAUCGAUUAUUUCCUUUUUUUAUU----- (((((((((((((..........))))).......(((((....)))))((((.......))))))))))))....................----- ( -26.60, z-score = -1.21, R) >droYak2.chr2R 6462705 92 - 21139217 GACUCCGUGGCGCAACGG-UAGCGCGUCCGACUCCAGAUCGGAAGGUUGCGUGUUCAAAUCACGUCGGGGUCAAUUGAUGAGUUAUAUUUUUA---- (((((....((((.....-..))))....((((((.((((....))))(((((.......))))).)))))).......))))).........---- ( -30.70, z-score = -1.52, R) >droSec1.super_1 12034769 92 + 14215200 GACUCCGUGGCGCAACGG-UAGCGCGUCCGACUCCAGAUCGGAAGGUUGCGUGUUCAAAUCACGUCGGGGUCAC--AAUGCAAGUAAUAUUUACU-- (.((((((......))))-.)))((((..((((((.((((....))))(((((.......))))).))))))..--.)))).((((.....))))-- ( -29.00, z-score = -1.12, R) >droSim1.chr2R 13231438 90 + 19596830 GACUCCGUGGCGCAACGG-UAGCGCGUCCGACUCCAGAUCGGAAGGUUGCGUGUUCAAAUCACGUCGGGGUCAAUGCAAUAGACUUUUUUU------ (.((((((......))))-.)))((((..((((((.((((....))))(((((.......))))).)))))).))))..............------ ( -29.40, z-score = -1.13, R) >consensus GACUCCGUGGCGCAACGG_UAGCGCGUCUGACUCCAGAUCAGAAGGUUGCGUGUUCAAAUCACGUCGGGGUCAAU_AAUGUGAUUUAUUUUU_____ (((((((..((((........)))).((((((....).))))).....(((((.......))))))))))))......................... (-28.29 = -27.15 + -1.13)


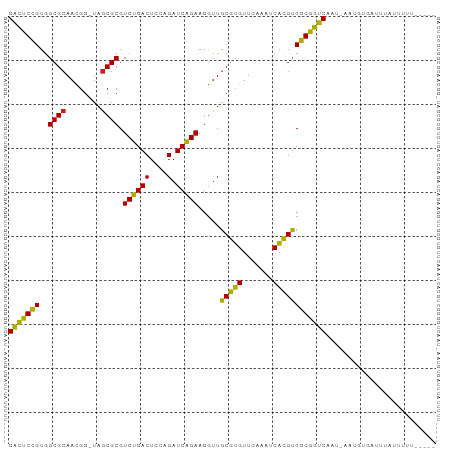
| Location | 14,527,061 – 14,527,150 |
|---|---|
| Length | 89 |
| Sequences | 11 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 80.00 |
| Shannon entropy | 0.44020 |
| G+C content | 0.52276 |
| Mean single sequence MFE | -21.87 |
| Consensus MFE | -22.24 |
| Energy contribution | -21.13 |
| Covariance contribution | -1.11 |
| Combinations/Pair | 1.30 |
| Mean z-score | -0.36 |
| Structure conservation index | 1.02 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.752322 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 14527061 89 - 21146708 ------AAAAAACAUCAAUUGC-UUGACCCCGACGUGAUUUGAACACGCAACCUUCCGAUCUGGAGUCGGACGCGCUA-CCGUUGCGCCACGGAGUC ------................-..(((.(((.((((.......))))......((((((.....)))))).((((..-.....))))..))).))) ( -22.50, z-score = -0.37, R) >anoGam1.chr2R 5380048 96 + 62725911 AAAGGAAUUGCACAUCAUAUUUUUUGACCCCGACGUGAUUUGAACACGCAACCUUCUGAUCUGGAGUCAGACGCGCUA-CCGUUGCGCCACGGAGUC .(((((((..........)))))))(((.(((.((((.......))))......((((((.....)))))).((((..-.....))))..))).))) ( -21.10, z-score = 0.24, R) >droGri2.scaffold_15245 13598005 90 - 18325388 -----AAAAAUCACUCGAGCU-CUUGACCCCGACGUGAAUCGAACACGCAACCUUCUGAUCUGGAGUCAGACGCGCUA-CCGUUGCGCCACGGAGUC -----................-...(((.(((.((((.......))))......((((((.....)))))).((((..-.....))))..))).))) ( -21.00, z-score = 0.34, R) >droMoj3.scaffold_6496 15789347 88 - 26866924 -----AAUAAUUUGUCACAUU---UGACCCCGACGUGAAUCGAACACGCAACCUUCUGAUCUGGAGUCAGACGCGCUA-CCGUUGCGCCACGGAGUC -----................---.(((.(((.((((.......))))......((((((.....)))))).((((..-.....))))..))).))) ( -21.00, z-score = -0.19, R) >droVir3.scaffold_12875 9473643 84 + 20611582 ------------AAAAACCUCGAUUGACCCCGACGUGAAUCGAACACGCAACCUUCUGAUCUGGAGUCAGACGCGCUA-CCGUUGCGCCACGGAGUC ------------.......((((((.((......)).))))))........((.((((((.....)))))).((((..-.....))))...)).... ( -21.10, z-score = -0.48, R) >droWil1.scaffold_180700 1819304 88 + 6630534 --------AAGAAAUCCCAAUCGUUGACCCCGACGUGAUUUGAACACGCAACCUUCUGAUCUGGAGUCAGACGCGCUA-CCGUUGCGCCACGGAGUC --------.................(((.(((.((((.......))))......((((((.....)))))).((((..-.....))))..))).))) ( -21.00, z-score = -0.09, R) >droPer1.super_31 71934 91 - 935084 -----UUUAAAAAAAUUAGUCUAUUGACCCCGACGUGAUUUGAACACGCAACCUUCUGAUCUGGAGUCAGACGCGCUA-CCGUUGCGCCACGGAGUC -----....................(((.(((.((((.......))))......((((((.....)))))).((((..-.....))))..))).))) ( -21.00, z-score = -0.73, R) >droAna3.scaffold_13266 17227175 92 + 19884421 -----AAUAAAAAAAGGAAAUAAUCGAUCCUGCCAGGAGUCGAACCUGGAAUCUUCUGAUCCGUAGUCAGACGCGUUAUCCAUUGCGCCACAGGACC -----..........((......(((((((.....))).)))).((((......((((((.....)))))).((((........))))..)))).)) ( -23.50, z-score = -1.59, R) >droYak2.chr2R 6462705 92 + 21139217 ----UAAAAAUAUAACUCAUCAAUUGACCCCGACGUGAUUUGAACACGCAACCUUCCGAUCUGGAGUCGGACGCGCUA-CCGUUGCGCCACGGAGUC ----.....................(((.(((.((((.......))))......((((((.....)))))).((((..-.....))))..))).))) ( -22.50, z-score = -0.86, R) >droSec1.super_1 12034769 92 - 14215200 --AGUAAAUAUUACUUGCAUU--GUGACCCCGACGUGAUUUGAACACGCAACCUUCCGAUCUGGAGUCGGACGCGCUA-CCGUUGCGCCACGGAGUC --.((((.......))))...--..(((.(((.((((.......))))......((((((.....)))))).((((..-.....))))..))).))) ( -23.40, z-score = -0.13, R) >droSim1.chr2R 13231438 90 - 19596830 ------AAAAAAAGUCUAUUGCAUUGACCCCGACGUGAUUUGAACACGCAACCUUCCGAUCUGGAGUCGGACGCGCUA-CCGUUGCGCCACGGAGUC ------...................(((.(((.((((.......))))......((((((.....)))))).((((..-.....))))..))).))) ( -22.50, z-score = -0.10, R) >consensus _____AAAAAAAAAUCAAAUU_AUUGACCCCGACGUGAUUUGAACACGCAACCUUCUGAUCUGGAGUCAGACGCGCUA_CCGUUGCGCCACGGAGUC .........................(((.(((.((((.......))))......((((((.....)))))).((((........))))..))).))) (-22.24 = -21.13 + -1.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:32:33 2011