
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 3,220,340 – 3,220,456 |
| Length | 116 |
| Max. P | 0.500000 |

| Location | 3,220,340 – 3,220,456 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 78.84 |
| Shannon entropy | 0.39007 |
| G+C content | 0.47860 |
| Mean single sequence MFE | -31.39 |
| Consensus MFE | -19.22 |
| Energy contribution | -18.77 |
| Covariance contribution | -0.45 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr2L 3220340 116 + 23011544 GAAUGCGGAUAGUGGGACAGUUUGUCAAGGUUUUUAUGUGGAAGUGAAAAACCGAAACGGCGACGGAUAUUUGUGUCCAGUUCAGCCGACGAGAAAGC-CCAUGGCUCAUUGAAGCC ...........(((((....((((((..(((((((((......))).)))))).....(((((((((((....))))).)))..)))))))))....)-))))((((......)))) ( -34.70, z-score = -1.66, R) >droEre2.scaffold_4929 3257422 100 + 26641161 GUAUGCGAAUAGUAGGACAGUUUGUCAAGGUUUUUAUGUGGAAGUGAAAAACCGAAGCGGC----------------CAGUUCAGCCGACGAUAAAGU-CCACGGCUCAUUGAAGCC ...(((.....)))((((..((((((..(((((((((......))).))))))...(((((----------------.......)))).)))))))))-))..((((......)))) ( -26.20, z-score = -1.30, R) >droYak2.chr2L 3209434 116 + 22324452 GAAUGAGAAUAGUAGGACAGUUUGUCAGGGUUUUUAUGUGGCAGUGAAAAACCGAAACGGCGACGGACAUUGGUGUCCAGUUCAGCCGACGAGAAAGU-CUACGGCUCAUUGAAGCC .((((((....(((((....((((((.((.(((((((......))))))).)).....(((((((((((....))))).)))..)))))))))....)-))))..))))))...... ( -37.70, z-score = -2.76, R) >droSec1.super_5 1369895 116 + 5866729 GAAUGCGGAUAGUGGGACAGUUUGUCAAGGUUUUUAUGUGGAAGUGAAAAACCGAAACGGCGACGGAUAUUUGUGUCCAGUUCAGCCGACGAGAAAGC-CCAUGGCUCAUUGAAGUC .((((.((...(((((....((((((..(((((((((......))).)))))).....(((((((((((....))))).)))..)))))))))....)-))))..))))))...... ( -34.30, z-score = -1.67, R) >droSim1.chr2L 3159479 116 + 22036055 GAAUGCGGAUAGUGGGACAGUUUGUCAAGGUUUUUAUGUGGAAGUGAAAAACCGAAACGGCGACGGAUAUUUGUGUCCAGUUCAGCCGACGAGAAAGC-CCAUGGCUCAUUGAAGCC ...........(((((....((((((..(((((((((......))).)))))).....(((((((((((....))))).)))..)))))))))....)-))))((((......)))) ( -34.70, z-score = -1.66, R) >dp4.chr4_group4 1295841 96 + 6586962 GGAUGGGGAAAGCAGAACAGUUUGUCCCAGUUUUUAU-----AAAGAGAAACCGAUGCGGCA----------------GGAUAUUUGGGUGGGAGAGUUCUAUGGCUCAUUGAGGCC ...((((..((((......))))..))))((((((..-----....))))))......(((.----------------.......(((((.(.((....)).).))))).....))) ( -20.72, z-score = 0.73, R) >consensus GAAUGCGGAUAGUAGGACAGUUUGUCAAGGUUUUUAUGUGGAAGUGAAAAACCGAAACGGCGACGGAUAUUUGUGUCCAGUUCAGCCGACGAGAAAGC_CCAUGGCUCAUUGAAGCC ...........(((((....((((((..(((((((...........))))))).....(((...(((((....)))))......)))))))))....).))))((((......)))) (-19.22 = -18.77 + -0.45)
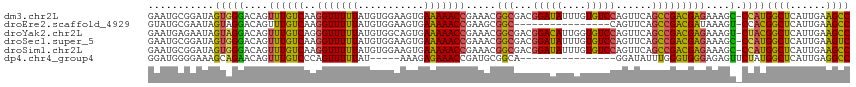

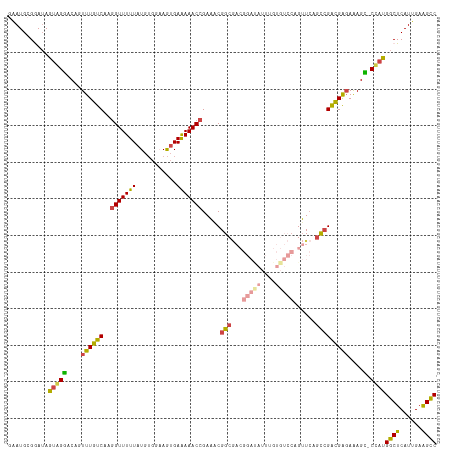
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:13:19 2011