
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 14,223,012 – 14,223,102 |
| Length | 90 |
| Max. P | 0.894481 |

| Location | 14,223,012 – 14,223,102 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 74.09 |
| Shannon entropy | 0.45453 |
| G+C content | 0.58029 |
| Mean single sequence MFE | -28.52 |
| Consensus MFE | -19.44 |
| Energy contribution | -19.25 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.894481 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr2R 14223012 90 + 21146708 AAGGUGAACUGAUGGCGUGUGGUUUUUGCCGCCGCUGCGGUUUCAAUUCAAGGGACCGUCCAUACGUCCAUCCAUUCAGCUACCCCCCAC--------------------- .((.((((..((((((((((((.....((....)).(((((..(.......)..)))))))))))).)))))..)))).)).........--------------------- ( -33.40, z-score = -2.87, R) >dp4.chr3 11499600 106 + 19779522 UGGGGGAACUGAUGGCGUGUGGUUUUUGCCGCCGUUGCGGUUUCAAUUCAAGUGACCGUCCC-----GUCCCCUUGUCGCGUCCCGUCACGUCCAUGCCACUGCCACUGCC (((((((((((.(((((.(((((....))))))))))))))))).......(((((.(..((-----(.(.....).)).)..).))))).))))................ ( -31.40, z-score = 0.94, R) >droPer1.super_4 6903425 105 + 7162766 UGGGGGAACUGAUGGCGUGUGGUUUUUGCCGCCGUUGCGGUUUCAAUUCAAGUGACCGUCC------GUCCCCUUGUCGCGUCCCGUCACGUCCAUGCCACUGCCACUGCC (((((((((((.(((((.(((((....))))))))))))))))).......(((((.(..(------((.........)))..).))))).))))................ ( -32.20, z-score = 0.64, R) >droAna3.scaffold_13266 15583360 80 - 19884421 AAGGUGAACUGAUGGCGUGUGGUUUUUGCCGCCGCUGCGGUUUCAAUUCAAGUGACCGUCC-------------CCCUGCCUCCCUCCUCGCC------------------ ..(((((......((((.(((((....)))))))))((((((.(.......).))))))..-------------..............)))))------------------ ( -24.30, z-score = -1.52, R) >droEre2.scaffold_4845 8444089 77 + 22589142 AAGGUGAACUGAUGGCGUGUGGUUUUUGCCGCCGCUGCGGUUUCAAUUCAAGUGACCGUCC------------AUCCAGCCCCCCCACC---------------------- ..((((..(((((((((.(((((....)))))))).((((((.(.......).)))))).)------------)).)))......))))---------------------- ( -23.90, z-score = -1.25, R) >droYak2.chr2R 6181108 93 - 21139217 AAGGUGAACUGAUGGCGUGUGGUUUUUGCCGCCGCUGCGGUUUCAAUUCAAGUGACCGUCCAUACGUCCAUCCAUUCAGCUCCCCUCCCCUCC------------------ .((.((((..((((((((((((.....((....)).((((((.(.......).))))))))))))).)))))..)))).))............------------------ ( -29.50, z-score = -2.91, R) >droSec1.super_1 11712353 84 + 14215200 AAGGUGAACUGAUGGCGUGUGGUUUUUGCCGCCGCUGCGGUUUCAAUUCAAGUGACCGUCC--------AUCCAUUCAGCUCUCCCCCCACC------------------- ..((.((.((((.((((.(((((....)))))))))((((((.(.......).))))))..--------......)))).)).)).......------------------- ( -26.90, z-score = -2.20, R) >droSim1.chr2R 12960341 82 + 19596830 AAGGUGAACUGAUGGCGUGUGGUUUUUGCCGCCGCUGCGGUUUCAAUUCAAGUGACCGUCC--------AUCCAUUCAGCUCCCCCCACC--------------------- .((.((((..(((((((.(((((....)))))))).((((((.(.......).)))))).)--------)))..)))).)).........--------------------- ( -26.60, z-score = -2.07, R) >consensus AAGGUGAACUGAUGGCGUGUGGUUUUUGCCGCCGCUGCGGUUUCAAUUCAAGUGACCGUCC________AUCCAUUCAGCUCCCCCCCCC_____________________ ..((.........((((.(((((....)))))))))(((((..(.......)..)))))........................)).......................... (-19.44 = -19.25 + -0.19)

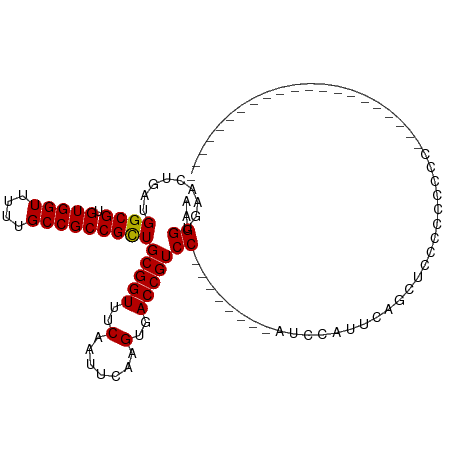
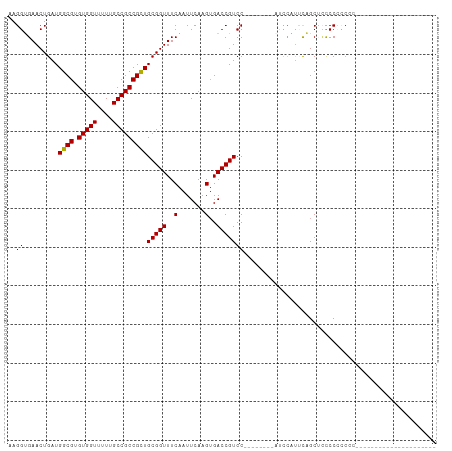
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:31:53 2011