
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 13,849,566 – 13,849,683 |
| Length | 117 |
| Max. P | 0.568113 |

| Location | 13,849,566 – 13,849,683 |
|---|---|
| Length | 117 |
| Sequences | 8 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.11 |
| Shannon entropy | 0.35904 |
| G+C content | 0.51762 |
| Mean single sequence MFE | -39.29 |
| Consensus MFE | -22.76 |
| Energy contribution | -23.61 |
| Covariance contribution | 0.85 |
| Combinations/Pair | 1.34 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.568113 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
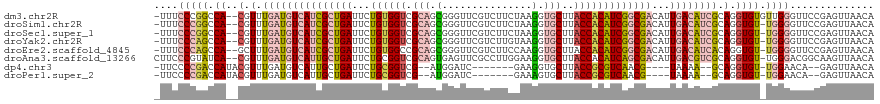
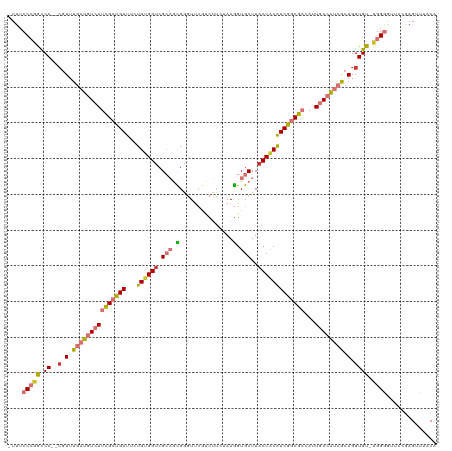
>dm3.chr2R 13849566 117 + 21146708 -UUUCCCGGCCA--CGUUUGAUGUCAUCGCUGAUUCUGUGGUCGCAGCGGGUUCGUCUUCUAAGGUGCUUACCACAUCGGCGACAUUGACAUCGCAGGUGUGUUGGGUUCCGAGUUAACA -...((((((((--(.(.(((((((((((((((...((((((.(((.((((.......)))..).)))..)))))))))))))...)))))))).).))).))))))............. ( -45.10, z-score = -2.68, R) >droSim1.chr2R 12585321 116 + 19596830 -UUUCCCGGCCA--CGUUUGAUGUCAUCGCUGAUUCUGUGGUCGCAGCGGGUUCGUCUUCUAAGGUGCUUACCACAUCGGCGACAUUGACAUCGCAGGUGU-UGGGGUUCCGAGUUAACA -..(((((((.(--(.(.(((((((((((((((...((((((.(((.((((.......)))..).)))..)))))))))))))...)))))))).).))))-)))))............. ( -42.90, z-score = -2.09, R) >droSec1.super_1 11344129 116 + 14215200 -UUUCCCGGCCA--CGUUUGAUGUCAUCGCUGAUUCUGUGGUCGCAGCGGGUUCGUCUUCUAAGGUGCUUACCACAUCGGCGACAUUGACAUCGCAGGUGU-UGGGGUUCCGAGUUAACA -..(((((((.(--(.(.(((((((((((((((...((((((.(((.((((.......)))..).)))..)))))))))))))...)))))))).).))))-)))))............. ( -42.90, z-score = -2.09, R) >droYak2.chr2R 5808233 116 - 21139217 -UUUCCCAGCCA--CGUUUGAUGUCAUCGCUGAUUCUGUGGUCGCAGCGGGUUCGUCUUGUAAGGUGCUUACCACAUCGGCGACAUUGACAUCGCAGGUGU-UGGGGUUCCGAGUUAACA -..(((((((.(--(.(.(((((((((((((((...(((((((((.(((((.....)))))...)))...)))))))))))))...)))))))).).))))-)))))............. ( -45.00, z-score = -2.78, R) >droEre2.scaffold_4845 8078201 116 + 22589142 -UUUCCCAGCCA--GCUUUGAUGUCAUCGCUGAUUCUGUGGCCGCAGCGGGUUCGUCUUCCAAGGUGCUUACCACAUCGGCGACAUUGACAUCACAGGUGU-UGGGGUUCCGAGUUAACA -..(((((((.(--.((.(((((((((((((((...(((((..(((.(.((........))..).)))...))))))))))))...)))))))).)).)))-)))))............. ( -44.80, z-score = -2.77, R) >droAna3.scaffold_13266 9543861 117 - 19884421 CUUCCCGUAUCA--CGUUUGAUGUCAUUGCUGAUUCUGCGGUCGCAGUGAGUUCGCCUUGGAAGGUGCUUACCACAUCAGCGACAUUGACGUCGCAGGUGU-UGGGACGGCAAGUUAACA ..(((((...((--(.(.((((((((((((((((..(((....)))(((.((.(((((....)))))...)))))))))))))...)))))))).).))).-)))))............. ( -41.90, z-score = -1.68, R) >dp4.chr3 6313701 101 + 19779522 -UUCCCCGACCAUACGUUUGAUGUCAUUGCUGAUUCUGCGGUCG--AUGGAUC-------GAAGGUGCUUACCGCGUCAACG----UAAAA--GCAGGUGU-UGGAACA--GAGUUAACA -((((...(((..((.((((((.(((((((((......))).))--)))))))-------))).))((((...(((....))----)..))--)).)))..-.))))..--......... ( -26.90, z-score = -0.02, R) >droPer1.super_2 6528437 101 + 9036312 -UUCCCCGACCAUACGUUUGAUGUCAUUGCUGAUUCUGCGGUCG--AUGGAUC-------GAAAGUGCUUACCGCGUCAACG----UAAAA--GCAGGUGU-UGGAACA--GAGUUAACA -((((...(((..((.((((((.(((((((((......))).))--)))))))-------))).))((((...(((....))----)..))--)).)))..-.))))..--......... ( -24.80, z-score = 0.27, R) >consensus _UUUCCCGGCCA__CGUUUGAUGUCAUCGCUGAUUCUGUGGUCGCAGCGGGUUCGUCUUCUAAGGUGCUUACCACAUCGGCGACAUUGACAUCGCAGGUGU_UGGGGUUCCGAGUUAACA ....(((((((.......(((((((((((((((...((((((.(((.(...............).)))..)))))))))))))...))))))))..)))...)))).............. (-22.76 = -23.61 + 0.85)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:31:02 2011